HLA DR Genome Editing with TALENs in Human iPSCs Produced Immune-Tolerant Dendritic Cells
- PMID: 34093711
- PMCID: PMC8163544
- DOI: 10.1155/2021/8873383
HLA DR Genome Editing with TALENs in Human iPSCs Produced Immune-Tolerant Dendritic Cells
Abstract
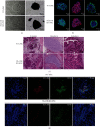
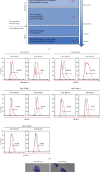
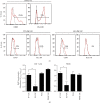
Although human induced pluripotent stem cells (iPSCs) can serve as a universal cell source for regenerative medicine, the use of iPSCs in clinical applications is limited by prohibitive costs and prolonged generation time. Moreover, allogeneic iPSC transplantation requires preclusion of mismatches between the donor and recipient human leukocyte antigen (HLA). We, therefore, generated universally compatible immune nonresponsive human iPSCs by gene editing. Transcription activator-like effector nucleases (TALENs) were designed for selective elimination of HLA DR expression. The engineered nucleases completely disrupted the expression of HLA DR on human dermal fibroblast cells (HDF) that did not express HLA DR even after stimulation with IFN-γ. Teratomas formed by HLA DR knockout iPSCs did not express HLA DR, and dendritic cells differentiated from HLA DR knockout iPSCs reduced CD4+ T cell activation. These engineered iPSCs might provide a novel translational approach to treat multiple recipients from a limited number of cell donors.
Copyright © 2021 Yoo-Wook Kwon et al.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Generation of hypoimmunogenic induced pluripotent stem cells by CRISPR-Cas9 system and detailed evaluation for clinical application.Mol Ther Methods Clin Dev. 2022 May 29;26:15-25. doi: 10.1016/j.omtm.2022.05.010. eCollection 2022 Sep 8. Mol Ther Methods Clin Dev. 2022. PMID: 35755947 Free PMC article.
-
CRISPR/Cas9 Ribonucleoprotein Complex-Mediated Efficient B2M Knockout in Human Induced Pluripotent Stem Cells (iPSCs).Methods Mol Biol. 2022;2454:607-624. doi: 10.1007/7651_2021_352. Methods Mol Biol. 2022. PMID: 33945142
-
Targeted Disruption of HLA Genes via CRISPR-Cas9 Generates iPSCs with Enhanced Immune Compatibility.Cell Stem Cell. 2019 Apr 4;24(4):566-578.e7. doi: 10.1016/j.stem.2019.02.005. Epub 2019 Mar 7. Cell Stem Cell. 2019. PMID: 30853558
-
Current status and future perspectives of HLA-edited induced pluripotent stem cells.Inflamm Regen. 2020 Oct 1;40:23. doi: 10.1186/s41232-020-00132-9. eCollection 2020. Inflamm Regen. 2020. PMID: 33014207 Free PMC article. Review.
-
Strategies for immune regulation in iPS cell-based cardiac regenerative medicine.Inflamm Regen. 2020 Sep 29;40:36. doi: 10.1186/s41232-020-00145-4. eCollection 2020. Inflamm Regen. 2020. PMID: 33005258 Free PMC article. Review.
Cited by
-
Reprogramming the tumor microenvironment by genome editing for precision cancer therapy.Mol Cancer. 2022 Apr 11;21(1):98. doi: 10.1186/s12943-022-01561-5. Mol Cancer. 2022. PMID: 35410257 Free PMC article. Review.
-
Status and prospects for the development of regenerative therapies for corneal and ocular diseases.Regen Ther. 2024 Sep 17;26:819-825. doi: 10.1016/j.reth.2024.09.001. eCollection 2024 Jun. Regen Ther. 2024. PMID: 39329098 Free PMC article. Review.
-
CAR-neutrophil mediated delivery of tumor-microenvironment responsive nanodrugs for glioblastoma chemo-immunotherapy.Nat Commun. 2023 Apr 20;14(1):2266. doi: 10.1038/s41467-023-37872-4. Nat Commun. 2023. PMID: 37080958 Free PMC article.
-
Emerging strategies for treating autoimmune disease with genetically modified dendritic cells.Cell Commun Signal. 2024 May 7;22(1):262. doi: 10.1186/s12964-024-01641-7. Cell Commun Signal. 2024. PMID: 38715122 Free PMC article. Review.
-
Managing allorejection in off-the-shelf CAR-engineered cell therapies.Mol Ther. 2025 Jun 4;33(6):2368-2390. doi: 10.1016/j.ymthe.2024.11.035. Epub 2024 Nov 26. Mol Ther. 2025. PMID: 39600090 Review.
References
LinkOut - more resources
Full Text Sources
Research Materials

