Bioinformatics Analysis of a Prognostic miRNA Signature and Potential Key Genes in Pancreatic Cancer
- PMID: 34094925
- PMCID: PMC8174116
- DOI: 10.3389/fonc.2021.641289
Bioinformatics Analysis of a Prognostic miRNA Signature and Potential Key Genes in Pancreatic Cancer
Abstract
Background: In this study, miRNAs and their critical target genes related to the prognosis of pancreatic cancer were screened based on bioinformatics analysis to provide targets for the prognosis and treatment of pancreatic cancer.
Methods: R software was used to screen differentially expressed miRNAs (DEMs) and genes (DEGs) downloaded from The Cancer Genome Atlas (TCGA) and Gene Expression Omnibus (GEO) databases, respectively. A miRNA Cox proportional hazards regression model was constructed based on the miRNAs, and a miRNA prognostic model was generated. The target genes of the prognostic miRNAs were predicted using TargetScan and miRDB and then intersected with the DEGs to obtain common genes. The functions of the common genes were subjected to Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses. A protein-protein interaction (PPI) network of the common genes was constructed with the STRING database and visualized with Cytoscape software. Key genes were also screened with the MCODE and cytoHubba plug-ins of Cytoscape. Finally, a prognostic model formed by the key gene was also established to help evaluate the reliability of this screening process.
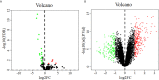
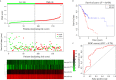
Results: A prognostic model containing four downregulated miRNAs (hsa-mir-424, hsa-mir-3613, hsa-mir-4772 and hsa-mir-126) related to the prognosis of pancreatic cancer was constructed. A total of 118 common genes were enriched in two KEGG pathways and 33 GO functional annotations, including extracellular matrix (ECM)-receptor interaction and cell adhesion. Nine key genes related to pancreatic cancer were also obtained: MMP14, ITGA2, THBS2, COL1A1, COL3A1, COL11A1, COL6A3, COL12A1 and COL5A2. The prognostic model formed by nine key genes also possessed good prognostic ability.
Conclusions: The prognostic model consisting of four miRNAs can reliably predict the prognosis of patients with pancreatic cancer. In addition, the screened nine key genes, which can also form a reliable prognostic model, are significantly related to the occurrence and development of pancreatic cancer. Among them, one novel miRNA (hsa-mir-4772) and two novel genes (COL12A1 and COL5A2) associated with pancreatic cancer have great potential to be used as prognostic factors and therapeutic targets for this tumor.
Keywords: Gene Expression Omnibus; The Cancer Genome Atlas; biomarkers; miRNAs; pancreatic cancer; target genes.
Copyright © 2021 Chen, Gao, Yu, Qu, Xiao and Huang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Identifying MMP14 and COL12A1 as a potential combination of prognostic biomarkers in pancreatic ductal adenocarcinoma using integrated bioinformatics analysis.PeerJ. 2020 Nov 23;8:e10419. doi: 10.7717/peerj.10419. eCollection 2020. PeerJ. 2020. PMID: 33282565 Free PMC article.
-
Identification of Prognostic miRNA Signature and Lymph Node Metastasis-Related Key Genes in Cervical Cancer.Front Pharmacol. 2020 May 8;11:544. doi: 10.3389/fphar.2020.00544. eCollection 2020. Front Pharmacol. 2020. PMID: 32457603 Free PMC article.
-
Bioinformatics Analysis of Prognostic miRNA Signature and Potential Critical Genes in Colon Cancer.Front Genet. 2020 Jun 9;11:478. doi: 10.3389/fgene.2020.00478. eCollection 2020. Front Genet. 2020. PMID: 32582275 Free PMC article.
-
Identification of Differentially Expressed Genes and miRNAs Associated with Esophageal Squamous Cell Carcinoma by Integrated Analysis of Microarray Data.Biomed Res Int. 2020 Jul 1;2020:1980921. doi: 10.1155/2020/1980921. eCollection 2020. Biomed Res Int. 2020. PMID: 32714975 Free PMC article.
-
Integrated analysis of mRNA-single nucleotide polymorphism-microRNA interaction network to identify biomarkers associated with prostate cancer.Front Genet. 2022 Jul 25;13:922712. doi: 10.3389/fgene.2022.922712. eCollection 2022. Front Genet. 2022. PMID: 35957689 Free PMC article.
Cited by
-
Integrating a microRNA signature as a liquid biopsy-based tool for the early diagnosis and prediction of potential therapeutic targets in pancreatic cancer.Br J Cancer. 2024 Jan;130(1):125-134. doi: 10.1038/s41416-023-02488-4. Epub 2023 Nov 10. Br J Cancer. 2024. PMID: 37950093 Free PMC article.
-
Exposure to trace levels of metals and fluoroquinolones increases inflammation and tumorigenesis risk of zebrafish embryos.Environ Sci Ecotechnol. 2022 Feb 25;10:100162. doi: 10.1016/j.ese.2022.100162. eCollection 2022 Apr. Environ Sci Ecotechnol. 2022. PMID: 36159734 Free PMC article.
-
Ultrasound-targeted microbubble destruction-mediated silencing of FBXO11 suppresses development of pancreatic cancer.Hum Cell. 2022 Jul;35(4):1174-1191. doi: 10.1007/s13577-022-00700-w. Epub 2022 Apr 18. Hum Cell. 2022. PMID: 35437704
-
A nine-gene signature as prognostic biomarker in gastric cancer by bioinformatics analysis.Clin Transl Oncol. 2023 Nov;25(11):3296-3306. doi: 10.1007/s12094-023-03180-y. Epub 2023 Apr 11. Clin Transl Oncol. 2023. PMID: 37041435
-
Role of Some microRNA/ADAM Proteins Axes in Gastrointestinal Cancers as a Novel Biomarkers and Potential Therapeutic Targets-A Review.Curr Issues Mol Biol. 2023 Apr 3;45(4):2917-2936. doi: 10.3390/cimb45040191. Curr Issues Mol Biol. 2023. PMID: 37185715 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

