Comprehensive Phosphoproteomic Analysis of Nostoc flagelliforme in Response to Dehydration Provides Insights into Plant ROS Signaling Transduction
- PMID: 34095650
- PMCID: PMC8173544
- DOI: 10.1021/acsomega.0c06111
Comprehensive Phosphoproteomic Analysis of Nostoc flagelliforme in Response to Dehydration Provides Insights into Plant ROS Signaling Transduction
Abstract
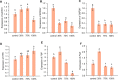
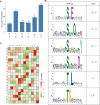
Terrestrial cyanobacteria, originated from aquatic cyanobacteria, exhibit a unique mechanism for drought adaptation during long-term evolution. To elucidate this diverse adaptive mechanism exhibited by terrestrial cyanobacteria from the post-translation modification aspect, we performed a global phosphoproteome analysis on the abundance of phosphoproteins in response to dehydration using Nostoc flagelliforme, a kind of terrestrial cyanobacteria having strong ecological adaptability to xeric environments. A total of 329 phosphopeptides from 271 phosphoproteins with 1168 phosphorylation sites were identified. Among these, 76 differentially expressed phosphorylated proteins (DEPPs) were identified for each dehydration treatment (30, 75, and 100% water loss), compared to control. The identified DEPPs were functionally categorized to be mainly involved in a two-component signaling pathway, photosynthesis, energy and carbohydrate metabolism, and an antioxidant system. We concluded that protein phosphorylation modifications related to the reactive oxygen species (ROS) signaling pathway might play an important role in coordinating enzyme activity involved in the antioxidant system in N. flagelliforme to adapt to dehydration stress. This study provides deep insights into the extensive modification of phosphorylation in terrestrial cyanobacteria using a phosphoproteomic approach, which may help to better understand the role of protein phosphorylation in key cellular mechanisms in terrestrial cyanobacteria in response to dehydration.
© 2021 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures





Similar articles
-
Effect of Drought Stress on Degradation and Remodeling of Membrane Lipids in Nostoc flagelliforme.Foods. 2022 Jun 18;11(12):1798. doi: 10.3390/foods11121798. Foods. 2022. PMID: 35741996 Free PMC article.
-
Global analysis of protein succinylation modification of Nostoc flagelliforme in response to dehydration.J Proteomics. 2021 Apr 15;237:104149. doi: 10.1016/j.jprot.2021.104149. Epub 2021 Feb 12. J Proteomics. 2021. PMID: 33588108
-
Enhancement of exopolysaccharides production and reactive oxygen species level of Nostoc flagelliforme in response to dehydration.Environ Sci Pollut Res Int. 2021 Jul;28(26):34300-34308. doi: 10.1007/s11356-021-13051-0. Epub 2021 Mar 1. Environ Sci Pollut Res Int. 2021. PMID: 33646551
-
The Enzyme Lysine Malonylation of Calvin Cycle and Gluconeogenesis Regulated Glycometabolism in Nostoc flagelliforme to Adapt to Drought Stress.Int J Mol Sci. 2023 May 8;24(9):8446. doi: 10.3390/ijms24098446. Int J Mol Sci. 2023. PMID: 37176152 Free PMC article.
-
Molecular Mechanisms of Nostoc flagelliforme Environmental Adaptation: A Comprehensive Review.Plants (Basel). 2025 May 22;14(11):1582. doi: 10.3390/plants14111582. Plants (Basel). 2025. PMID: 40508256 Free PMC article. Review.
Cited by
-
Reveal the kernel dehydration mechanisms in maize based on proteomic and metabolomic analysis.BMC Plant Biol. 2024 Jan 2;24(1):15. doi: 10.1186/s12870-023-04692-z. BMC Plant Biol. 2024. PMID: 38163910 Free PMC article.
-
Unveiling metabolic pathways involved in the extreme desiccation tolerance of an Atacama cyanobacterium.Sci Rep. 2023 Sep 22;13(1):15767. doi: 10.1038/s41598-023-41879-8. Sci Rep. 2023. PMID: 37737281 Free PMC article.
-
The regulatory impact of serine/threonine-specific protein phosphorylation among cyanobacteria.Front Plant Sci. 2025 Feb 12;16:1540914. doi: 10.3389/fpls.2025.1540914. eCollection 2025. Front Plant Sci. 2025. PMID: 40012730 Free PMC article. Review.
-
Proteome profiling reveals changes in energy metabolism, transport and antioxidation during drought stress in Nostoc flagelliforme.BMC Plant Biol. 2022 Apr 1;22(1):162. doi: 10.1186/s12870-022-03542-8. BMC Plant Biol. 2022. PMID: 35365086 Free PMC article.
-
Effect of Drought Stress on Degradation and Remodeling of Membrane Lipids in Nostoc flagelliforme.Foods. 2022 Jun 18;11(12):1798. doi: 10.3390/foods11121798. Foods. 2022. PMID: 35741996 Free PMC article.
References
-
- Rajeev L.; da Rocha U. N.; Klitgord N.; Luning E. G.; Fortney J.; Axen S. D.; Shih P. M.; Bouskill N. J.; Bowen B. P.; Kerfeld C. A.; Garcia-Pichel F.; Brodie E. L.; Northen T. R.; Mukhopadhyay A. Dynamic Cyanobacterial Response to Hydration and Dehydration in a Desert Biological Soil Crust. ISME J 2013, 7, 2178–2191. 10.1038/ismej.2013.83. - DOI - PMC - PubMed
-
- Qiu B.; Gao K. Daily Production and Photosynthetic Characteristics of Nostoc Flagelliforme Grown under Ambient and Elevated CO2 Conditions. J. Appl. Phycol. 2002, 14, 77–83. 10.1023/A:1019434414245. - DOI
-
- Gao K. Chinese Studies on the Edible Blue-Green Alga, Nostoc Flagelliforme: A Review. J. Appl. Phycol. 1998, 10, 37–49. 10.1023/A:1008014424247. - DOI
LinkOut - more resources
Full Text Sources

