Postnatal expression profiles of atypical cadherin FAT1 suggest its role in autism
- PMID: 34100899
- PMCID: PMC8214424
- DOI: 10.1242/bio.056457
Postnatal expression profiles of atypical cadherin FAT1 suggest its role in autism
Abstract
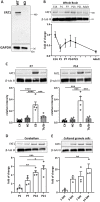
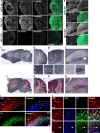
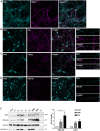
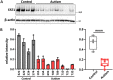
Genetic studies have linked FAT1 (FAT atypical cadherin 1) with autism spectrum disorder (ASD); however, the role that FAT1 plays in ASD remains unknown. In mice, the function of Fat1 has been primarily implicated in embryonic nervous system development with less known about its role in postnatal development. We show for the first time that FAT1 protein is expressed in mouse postnatal brains and is enriched in the cerebellum, where it localizes to granule neurons and Golgi cells in the granule layer, as well as inhibitory neurons in the molecular layer. Furthermore, subcellular characterization revealed FAT1 localization in neurites and soma of granule neurons, as well as being present in the synaptic plasma membrane and postsynaptic densities. Interestingly, FAT1 expression was decreased in induced pluripotent stem cell (iPSC)-derived neural precursor cells (NPCs) from individuals with ASD. These findings suggest a novel role for FAT1 in postnatal development and may be particularly important for cerebellum function. As the cerebellum is one of the vulnerable brain regions in ASD, our study warrants further investigation of FAT1 in the disease etiology.
Keywords: Autism; Cadherin; Cerebellum; FAT1; Granule cells; Neural precursor cells.
© 2021. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures
References
-
- Abou Jamra, R., Becker, T., Georgi, A., Feulner, T., Schumacher, J., Stromaier, J., Schirmbeck, F., Schulze, T. G., Propping, P., Rietschel, M.et al. (2008). Genetic variation of the FAT gene at 4q35 is associated with bipolar affective disorder. Mol. Psychiatry 13, 277-284. 10.1038/sj.mp.4002111 - DOI - PubMed
-
- American Psychiatric Association (2013). Diagnostic and Statistical Manual of Mental Disorders, 5th edn. Arlington, VA: American Psychiatric Publishing.
-
- Badouel, C., Zander, M. A., Liscio, N., Bagherie-Lachidan, M., Sopko, R., Coyaud, E., Raught, B., Miller, F. D. and McNeill, H. (2015). Fat1 interacts with Fat4 to regulate neural tube closure, neural progenitor proliferation and apical constriction during mouse brain development. Development 142, 2781-2791. 10.1242/dev.123539 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

