18S rRNA variability maps reveal three highly divergent, conserved motifs within Rotifera
- PMID: 34112085
- PMCID: PMC8194223
- DOI: 10.1186/s12862-021-01845-2
18S rRNA variability maps reveal three highly divergent, conserved motifs within Rotifera
Erratum in
-
Correction: 18S rRNA variability maps reveal three highly divergent, conserved motifs within Rotifera.BMC Ecol Evol. 2023 Jun 1;23(1):23. doi: 10.1186/s12862-023-02128-8. BMC Ecol Evol. 2023. PMID: 37264305 Free PMC article. No abstract available.
Abstract
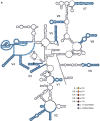
Background: 18S rRNA is a major component of the small subunit of the eukaryotic ribosome and an important phylogenetic marker for many groups, often to the point of being the only marker available for some. A core structure across eukaryotes exists for this molecule that can help to inform about its evolution in different groups. Using an alignment of 18S rDNA for Rotifera as traditionally recognized (=Bdelloidea, Monogononta, and Seisonacea, but not Acanthocephala), I fitted sequences for three exemplar species (Adineta vaga, Brachionus plicatilis, and Seison nebaliae, respectively) to the core structure and used these maps to reveal patterns of evolution for the remainder of this diverse group of microscopic animals.
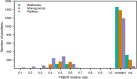
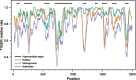
Results: The obtained variability maps of the 18S rRNA molecule revealed a pattern of high diversity among the three major rotifer clades coupled with strong conservation within each of bdelloids and monogononts. A majority of individual sites (ca. 60%) were constant even across rotifers as a whole with variable sites showing only intermediate rates of evolution. Although the three structural maps each showed good agreement with the inferred core structure for eukaryotic 18S rRNA and so were highly similar to one another at the secondary and tertiary levels, the overall pattern is of three highly distinct, but conserved motifs within the group at the primary sequence level. A novel finding was that of a variably expressed deletion at the 3' end of the V3 hypervariable region among some bdelloid species that occasionally extended into and included the pseudoknot structure following this region as well as the central "square" of the 18S rRNA molecule. Compared to other groups, levels of variation and rates of evolution for 18S rRNA in Rotifera roughly matched those for Gastropoda and Acanthocephala, despite increasing evidence for the latter being a clade within Rotifera.
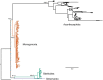
Conclusions: The lack of comparative data for comparable groups makes interpretation of the results (i.e., very low variation within each of the three major rotifer clades, but high variation between them) and their potential novelty difficult. However, these findings in combination with the high morphological diversity within rotifers potentially help to explain why no clear consensus has been reached to date with regard to the phylogenetic relationships among the major groups.
Keywords: Acanthocephala; Bdelloidea; Conservation; Deletion; Evolution; Hypervariable; Monogononta; Phylogeny; Rate of evolution; Seisonacea.
Conflict of interest statement
The author declares that he has no competing interests.
Figures




References
-
- Pleij CWA. Pseudoknots: a new motif in the RNA game. Trends Biochem Sci. 1990;15(4):143–147. - PubMed
-
- Ki J-S. Hypervariable regions (V1–V9) of the dinoflagellate 18S rRNA using a large dataset for marker considerations. J Appl Phycol. 2012;24(5):1035–1043.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
