Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states
- PMID: 34115987
- PMCID: PMC8782207
- DOI: 10.1016/j.ccell.2021.05.005
Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states
Abstract
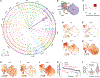
The underpinnings of cancer metastasis remain poorly understood, in part due to a lack of tools for probing their emergence at high resolution. Here we present macsGESTALT, an inducible CRISPR-Cas9-based lineage recorder with highly efficient single-cell capture of both transcriptional and phylogenetic information. Applying macsGESTALT to a mouse model of metastatic pancreatic cancer, we recover ∼380,000 CRISPR target sites and reconstruct dissemination of ∼28,000 single cells across multiple metastatic sites. We find that cells occupy a continuum of epithelial-to-mesenchymal transition (EMT) states. Metastatic potential peaks in rare, late-hybrid EMT states, which are aggressively selected from a predominately epithelial ancestral pool. The gene signatures of these late-hybrid EMT states are predictive of reduced survival in both human pancreatic and lung cancer patients, highlighting their relevance to clinical disease progression. Finally, we observe evidence for in vivo propagation of S100 family gene expression across clonally distinct metastatic subpopulations.
Keywords: CRISPR; EMT; S100; barcoding; epithelial-to-mesenchymal transition; evolving barcodes; lineage tracing; metastasis; phylogenetics; single cell.
Copyright © 2021 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declartion of interests The authors declare no competing interests.
Figures





Comment in
-
Lineage tracing reveals metastatic dynamics.Cancer Cell. 2021 Aug 9;39(8):1050-1052. doi: 10.1016/j.ccell.2021.06.005. Epub 2021 Jun 24. Cancer Cell. 2021. PMID: 34171265
References
-
- Aiello NM, Rhim AD, and Stanger BZ (2016). Orthotopic injection of pancreatic cancer cells. Cold Spring Harb. Protoc 2016, db.prot078360. - PubMed
-
- Beard C, Hochedlinger K, Plath K, Anton W, and Jaenisch R (2006). Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

