Sustained viremia suppression by SHIVSF162P3CN-recalled effector-memory CD8+ T cells after PD1-based vaccination
- PMID: 34125864
- PMCID: PMC8202916
- DOI: 10.1371/journal.ppat.1009647
Sustained viremia suppression by SHIVSF162P3CN-recalled effector-memory CD8+ T cells after PD1-based vaccination
Abstract
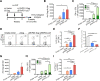
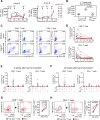
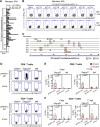
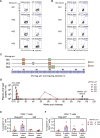
HIV-1 functional cure requires sustained viral suppression without antiretroviral therapy. While effector-memory CD8+ T lymphocytes are essential for viremia control, few vaccines elicit such cellular immunity that could be potently recalled upon viral infection. Here, we investigated a program death-1 (PD1)-based vaccine by fusion of simian immunodeficiency virus capsid antigen to soluble PD1. Homologous vaccinations suppressed setpoint viremia to undetectable levels in vaccinated macaques following a high-dose intravenous challenge by the pathogenic SHIVSF162P3CN. Poly-functional effector-memory CD8+ T cells were not only induced after vaccination, but were also recalled upon viral challenge for viremia control as determined by CD8 depletion. Vaccine-induced effector memory CD8+ subsets displayed high cytotoxicity-related genes by single-cell analysis. Vaccinees with sustained viremia suppression for over two years responded to boost vaccination without viral rebound. These results demonstrated that PD1-based vaccine-induced effector-memory CD8+ T cells were recalled by AIDS virus infection, providing a potential immunotherapy for functional cure.
Conflict of interest statement
I have read the journal’s policy and the authors of this manuscript have the following competing interests: Z.C. is a co-inventor on the PD1-based vaccine patent. However, ZC and all other authors do not have any competing interests or additional financial interests.
Figures


Similar articles
-
Containment of simian immunodeficiency virus infection in vaccinated macaques: correlation with the magnitude of virus-specific pre- and postchallenge CD4+ and CD8+ T cell responses.J Immunol. 2002 Nov 1;169(9):4778-87. doi: 10.4049/jimmunol.169.9.4778. J Immunol. 2002. PMID: 12391187
-
Sustained suppression of SHIV89.6P replication in macaques by vaccine-induced CD8+ memory T cells.AIDS. 2008 Sep 12;22(14):1739-48. doi: 10.1097/QAD.0b013e32830efdae. AIDS. 2008. PMID: 18753858
-
CD8 T Cells Show Protection against Highly Pathogenic Simian Immunodeficiency Virus (SIV) after Vaccination with SIV Gene-Expressing BCG Prime and Vaccinia Virus/Sendai Virus Vector Boosts.J Virol. 2021 Jan 28;95(4):e01718-20. doi: 10.1128/JVI.01718-20. Print 2021 Jan 28. J Virol. 2021. PMID: 33087465 Free PMC article.
-
Elevated expression levels of inhibitory receptor programmed death 1 on simian immunodeficiency virus-specific CD8 T cells during chronic infection but not after vaccination.J Virol. 2007 Jun;81(11):5819-28. doi: 10.1128/JVI.00024-07. Epub 2007 Mar 21. J Virol. 2007. PMID: 17376899 Free PMC article.
-
Vaccination of Macaques with DNA Followed by Adenoviral Vectors Encoding Simian Immunodeficiency Virus (SIV) Gag Alone Delays Infection by Repeated Mucosal Challenge with SIV.J Virol. 2019 Oct 15;93(21):e00606-19. doi: 10.1128/JVI.00606-19. Print 2019 Nov 1. J Virol. 2019. PMID: 31413132 Free PMC article.
Cited by
-
An open-label study on the safety and immunogenicity of a PD-1-enhanced DNA vaccine used as a T cell booster for COVID-19.EBioMedicine. 2025 May;115:105699. doi: 10.1016/j.ebiom.2025.105699. Epub 2025 Apr 16. EBioMedicine. 2025. PMID: 40245494 Free PMC article. Clinical Trial.
-
Grand Challenges on HIV/AIDS in China - The 5th Symposium, Yunnan 2024.Emerg Microbes Infect. 2025 Dec;14(1):2492208. doi: 10.1080/22221751.2025.2492208. Epub 2025 Apr 22. Emerg Microbes Infect. 2025. PMID: 40202047 Free PMC article.
-
Enhanced Cross-Reactive and Polyfunctional Effector-Memory T Cell Responses by ICVAX-a Human PD1-Based Bivalent HIV-1 Gag-p41 Mosaic DNA Vaccine.J Virol. 2022 Apr 13;96(7):e0216121. doi: 10.1128/jvi.02161-21. Epub 2022 Mar 17. J Virol. 2022. PMID: 35297660 Free PMC article.
References
-
- Sáez-Cirión A, Lacabaratz C, Lambotte O, Versmisse P, Urrutia A, Boufassa F, et al.. HIV controllers exhibit potent CD8 T cell capacity to suppress HIV infection ex vivo and peculiar cytotoxic T lymphocyte activation phenotype. Proc Natl Acad Sci U S A. 2007;104(16):6776–81. doi: 10.1073/pnas.0611244104 Proceedings of the National Academy of Sciences. - DOI - PMC - PubMed
-
- Koup RA, Safrit JT, Cao Y, Andrews CA, McLeod G, Borkowsky W, et al.. Temporal association of cellular immune responses with the initial control of viremia in primary human immunodeficiency virus type 1 syndrome. Journal of Virology. 1994;68(7):4650–5. doi: 10.1128/JVI.68.7.4650-4655.1994 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

