This is a preprint.
Type I Interferon Transcriptional Network Regulates Expression of Coinhibitory Receptors in Human T cells
- PMID: 34127967
- PMCID: PMC8202434
- DOI: 10.21203/rs.3.rs-133494/v1
Type I Interferon Transcriptional Network Regulates Expression of Coinhibitory Receptors in Human T cells
Update in
-
Type I interferon transcriptional network regulates expression of coinhibitory receptors in human T cells.Nat Immunol. 2022 Apr;23(4):632-642. doi: 10.1038/s41590-022-01152-y. Epub 2022 Mar 17. Nat Immunol. 2022. PMID: 35301508 Free PMC article.
Abstract
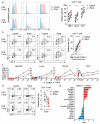
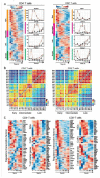
While inhibition of T cell co-inhibitory receptors has revolutionized cancer therapy, the mechanisms governing their expression on human T cells have not been elucidated. Type 1 interferon (IFN-I) modulates T cell immunity in viral infection, autoimmunity, and cancer, and may facilitate induction of T cell exhaustion in chronic viral infection. Here we show that IFN-I regulates co-inhibitory receptor expression on human T cells, inducing PD-1/TIM-3/LAG-3 while surprisingly inhibiting TIGIT expression. High-temporal-resolution mRNA profiling of IFN-I responses enabled the construction of dynamic transcriptional regulatory networks uncovering three temporal transcriptional waves. Perturbation of key transcription factors on human primary T cells revealed unique regulators that control expression of co-inhibitory receptors. We found that the dynamic IFN-I response in vitro closely mirrored T cell features with IFN-I linked acute SARS-CoV-2 infection in human, with high LAG3 and decreased TIGIT expression. Finally, our gene regulatory network identified SP140 as a key regulator for differential LAG3 and TIGIT expression, which were validated at the level of protein expression. The construction of IFN-I regulatory networks with identification of unique transcription factors controlling co-inhibitory receptor expression may provide targets for enhancement of immunotherapy in cancer, infectious diseases, and autoimmunity.
Figures



References
Publication types
Grants and funding
- R01 CA227473/CA/NCI NIH HHS/United States
- UL1 TR001863/TR/NCATS NIH HHS/United States
- R01 NS045937/NS/NINDS NIH HHS/United States
- P01 AI039671/AI/NIAID NIH HHS/United States
- UM1 HG009390/HG/NHGRI NIH HHS/United States
- R25 NS079193/NS/NINDS NIH HHS/United States
- P50 CA121974/CA/NCI NIH HHS/United States
- U19 AI089992/AI/NIAID NIH HHS/United States
- R01 AI144166/AI/NIAID NIH HHS/United States
- P01 AI073748/AI/NIAID NIH HHS/United States
- U01 HL145567/HL/NHLBI NIH HHS/United States
- R01 HL141852/HL/NHLBI NIH HHS/United States
- R01 NS030843/NS/NINDS NIH HHS/United States
- P01 AI056299/AI/NIAID NIH HHS/United States
- R01 HL127349/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

