Recent advances (2019-2021) of capillary electrophoresis-mass spectrometry for multilevel proteomics
- PMID: 34128246
- PMCID: PMC8671558
- DOI: 10.1002/mas.21714
Recent advances (2019-2021) of capillary electrophoresis-mass spectrometry for multilevel proteomics
Abstract
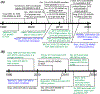
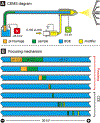
Multilevel proteomics aims to delineate proteins at the peptide (bottom-up proteomics), proteoform (top-down proteomics), and protein complex (native proteomics) levels. Capillary electrophoresis-mass spectrometry (CE-MS) can achieve highly efficient separation and highly sensitive detection of complex mixtures of peptides, proteoforms, and even protein complexes because of its substantial technical progress. CE-MS has become a valuable alternative to the routinely used liquid chromatography-mass spectrometry for multilevel proteomics. This review summarizes the most recent (2019-2021) advances of CE-MS for multilevel proteomics regarding technological progress and biological applications. We also provide brief perspectives on CE-MS for multilevel proteomics at the end, highlighting some future directions and potential challenges.
Keywords: capillary isoelectric focusing-mass spectrometry; capillary zone electrophoresis-mass spectrometry; multilevel proteomics; protein complex; proteoform.
© 2021 John Wiley & Sons Ltd.
Conflict of interest statement
Conflict of interest
The authors declared no conflict of interest related to this work.
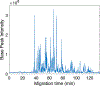
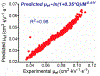
Figures
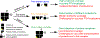

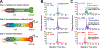

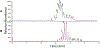

Similar articles
-
[Capillary electrophoresis-mass spectrometry and its application to proteomic analysis].Se Pu. 2020 Oct 8;38(10):1117-1124. doi: 10.3724/SP.J.1123.2020.03005. Se Pu. 2020. PMID: 34213108 Review. Chinese.
-
Capillary Electrophoresis-Mass Spectrometry for Top-Down Proteomics.Annu Rev Anal Chem (Palo Alto Calif). 2025 May;18(1):125-147. doi: 10.1146/annurev-anchem-071124-092242. Epub 2025 Jan 23. Annu Rev Anal Chem (Palo Alto Calif). 2025. PMID: 39847747 Review.
-
Top-Down Proteomics by Capillary Zone Electrophoresis-Tandem Mass Spectrometry for Large-Scale Characterization of Proteoforms in Complex Samples.Methods Mol Biol. 2022;2531:107-124. doi: 10.1007/978-1-0716-2493-7_8. Methods Mol Biol. 2022. PMID: 35941482
-
Recent trends of capillary electrophoresis-mass spectrometry in proteomics research.Mass Spectrom Rev. 2019 Nov;38(6):445-460. doi: 10.1002/mas.21599. Epub 2019 Aug 12. Mass Spectrom Rev. 2019. PMID: 31407381 Free PMC article. Review.
-
Large-Scale Qualitative and Quantitative Top-Down Proteomics Using Capillary Zone Electrophoresis-Electrospray Ionization-Tandem Mass Spectrometry with Nanograms of Proteome Samples.J Am Soc Mass Spectrom. 2019 Aug;30(8):1435-1445. doi: 10.1007/s13361-019-02167-w. Epub 2019 Apr 9. J Am Soc Mass Spectrom. 2019. PMID: 30972727 Free PMC article.
Cited by
-
Technological advances for analyzing the content of organ-on-a-chip by mass spectrometry.Front Bioeng Biotechnol. 2023 May 22;11:1197760. doi: 10.3389/fbioe.2023.1197760. eCollection 2023. Front Bioeng Biotechnol. 2023. PMID: 37284240 Free PMC article. Review.
-
Native Proteomics by Capillary Zone Electrophoresis-Mass Spectrometry.Angew Chem Int Ed Engl. 2024 Nov 25;63(48):e202408370. doi: 10.1002/anie.202408370. Epub 2024 Oct 24. Angew Chem Int Ed Engl. 2024. PMID: 39196601
-
Coupling High-Field Asymmetric Waveform Ion Mobility Spectrometry with Capillary Zone Electrophoresis-Tandem Mass Spectrometry for Top-Down Proteomics.Anal Chem. 2023 Jun 27;95(25):9497-9504. doi: 10.1021/acs.analchem.3c00551. Epub 2023 May 30. Anal Chem. 2023. PMID: 37254456 Free PMC article.
-
The role of protein corona in advancing plasma proteomics.Proteomics. 2025 Jan;25(1-2):e2400028. doi: 10.1002/pmic.202400028. Epub 2024 Sep 2. Proteomics. 2025. PMID: 39221533 Free PMC article. Review.
-
Evaluation of Machine Learning Models for Proteoform Retention and Migration Time Prediction in Top-Down Mass Spectrometry.J Proteome Res. 2022 Jul 1;21(7):1736-1747. doi: 10.1021/acs.jproteome.2c00124. Epub 2022 May 26. J Proteome Res. 2022. PMID: 35616364 Free PMC article.
References
-
- Aebersold R, Mann M. Mass-spectrometric exploration of proteome structure and function. Nature. 2016;537:347–55. - PubMed
-
- Akashi S, Suzuki K, Arai A, Yamada N, Suzuki E, Hirayama K, Nakamura S, Nishimura Y. Top-down analysis of basic proteins by microchip capillary electrophoresis mass spectrometry. Rapid Commun. Mass Spectrom. 2006;20:1932–8. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

