Orthobunyaviruses in the Caribbean: Melao and Oropouche virus infections in school children in Haiti in 2014
- PMID: 34133422
- PMCID: PMC8238191
- DOI: 10.1371/journal.pntd.0009494
Orthobunyaviruses in the Caribbean: Melao and Oropouche virus infections in school children in Haiti in 2014
Abstract
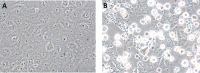
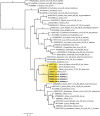
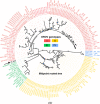
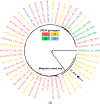
We report the identification of two orthobunyaviruses, Melao virus (MELV) and Oropouche virus (OROV), in plasma specimens from Haitian children with acute febrile illness who presented during outbreaks caused by alpha- and flaviviruses in 2014. Heretofore not described as a human pathogen, MELV was isolated in cell culture from the plasma of five case patients. OROV RNA was detected in the plasma of an additional child, using an unbiased sequencing approach, with phylogenetic inference suggesting a close relationship with strains from Brazil. Abdominal pain was reported by four case patients with MELV infections, with lymphadenopathy noted in two cases. Our findings document the occurrence of these orthobunyaviruses within the Caribbean region and highlight the critical importance of surveillance with viral genome sequence analyses to identify outbreaks caused by these and other emerging viruses.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




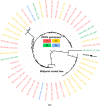
References
-
- https://talk.ictvonline.org/ictv-reports/ictv_online_report/negative-sen... (accessed 22 Feb. 2021).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

