Fusion gene map of acute leukemia revealed by transcriptome sequencing of a consecutive cohort of 1000 cases in a single center
- PMID: 34135310
- PMCID: PMC8209121
- DOI: 10.1038/s41408-021-00504-5
Fusion gene map of acute leukemia revealed by transcriptome sequencing of a consecutive cohort of 1000 cases in a single center
Abstract
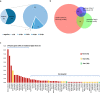
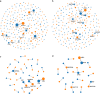
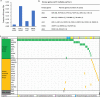
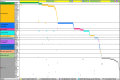
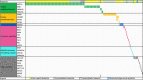
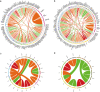
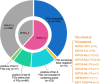
Fusion genes (FGs) are important genetic abnormalities in acute leukemias, but their variety and occurrence in acute leukemias remain to be systematically described. Whole transcriptome sequencing (WTS) provides a powerful tool for analyzing FGs. Here we report the FG map revealed by WTS in a consecutive cohort of 1000 acute leukemia cases in a single center, including 539 acute myeloid leukemia (AML), 437 acute lymphoblastic leukemia (ALL), and 24 mixed-phenotype acute leukemia (MPAL) patients. Bioinformatic analysis identified 792 high-confidence in-frame fusion events (296 distinct fusions) which were classified into four tiers. Tier A (pathogenic), B (likely pathogenic), and C (uncertain significance) FGs were identified in 61.8% cases of the total cohort (59.7% in AML, 64.5% in ALL, and 63.6% in MPAL). FGs involving protein kinase, transcription factor, and epigenetic genes were detected in 10.7%, 48.5%, and 15.1% cases, respectively. A considerable amount of novel FGs (82 in AML, 88 in B-ALL, 13 in T-ALL, and 9 in MPAL) was identified. This comprehensively described real map of FGs in acute leukemia revealed multiple FGs with clinical relevance that have not been previously recognized. WTS is a valuable tool and should be widely used in the routine diagnostic workup of acute leukemia.
Conflict of interest statement
The authors declare no competing interests.
Figures
Similar articles
-
Clinical application of whole transcriptome sequencing for the classification of patients with acute lymphoblastic leukemia.BMC Cancer. 2021 Aug 2;21(1):886. doi: 10.1186/s12885-021-08635-5. BMC Cancer. 2021. PMID: 34340673 Free PMC article.
-
The incidence, genetic characteristics, and prognosis of leukemia with concurrent pathogenic fusion genes: a series of 25 cases from a large cohort of leukemia patients.Cancer Gene Ther. 2020 Feb;27(1-2):89-97. doi: 10.1038/s41417-019-0147-1. Epub 2019 Oct 23. Cancer Gene Ther. 2020. PMID: 31645680
-
[Biological microchip for establishing the structure of fusion transcripts involving MLL in children with acute leukemia].Mol Biol (Mosk). 2016 Nov-Dec;50(6):968-977. doi: 10.7868/S0026898416060148. Mol Biol (Mosk). 2016. PMID: 28064313 Russian.
-
Single-cell RNA sequencing distinctly characterizes the wide heterogeneity in pediatric mixed phenotype acute leukemia.Genome Med. 2023 Oct 16;15(1):83. doi: 10.1186/s13073-023-01241-z. Genome Med. 2023. PMID: 37845689 Free PMC article.
-
Mixed Phenotype Acute Leukemia with t(12;17)(p13;q21)/TAF15-ZNF384 and Other Chromosome Abnormalities.Cytogenet Genome Res. 2016;149(3):165-170. doi: 10.1159/000448447. Epub 2016 Sep 9. Cytogenet Genome Res. 2016. PMID: 27607436 Review.
Cited by
-
Implementation of Whole-Genome and Transcriptome Sequencing Into Clinical Cancer Care.JCO Precis Oncol. 2022 Dec;6:e2200245. doi: 10.1200/PO.22.00245. JCO Precis Oncol. 2022. PMID: 36480778 Free PMC article. Review.
-
First report of B-lymphoblastic leukemia harboring LRRFIP1::FGFR1 fusion: expanding the clinical spectrum of myeloid/lymphoid neoplasms with tyrosine kinase gene fusions.Ann Hematol. 2025 Jul;104(7):3899-3902. doi: 10.1007/s00277-025-06481-0. Epub 2025 Jun 30. Ann Hematol. 2025. PMID: 40586868 Free PMC article.
-
Molecular characterization and biomarker identification in paediatric B-cell acute lymphoblastic leukaemia.J Cell Mol Med. 2024 Oct;28(19):e70126. doi: 10.1111/jcmm.70126. J Cell Mol Med. 2024. PMID: 39384181 Free PMC article.
-
Case Report: Identification of a novel LYN::LINC01900 transcript with promyelocytic phenotype and TP53 mutation in acute myeloid leukemia.Front Oncol. 2023 Dec 1;13:1322403. doi: 10.3389/fonc.2023.1322403. eCollection 2023. Front Oncol. 2023. PMID: 38107067 Free PMC article.
-
A novel gene fusion RUNX1/ZNF423 promotes leukemic relapse of NUP98-rearranged AML.Leukemia. 2023 Nov;37(11):2286-2291. doi: 10.1038/s41375-023-02024-6. Epub 2023 Sep 15. Leukemia. 2023. PMID: 37714925 No abstract available.
References
-
- Harris NL, Jaffe ES, Diebold J, Flandrin G, Muller-Hermelink HK, Vardiman J, et al. The World Health Organization classification of neoplastic diseases of the haematopoietic and lymphoid tissues: Report of the Clinical Advisory Committee Meeting, Airlie House, Virginia, November 1997. Histopathology. 2000;36:69–86. doi: 10.1046/j.1365-2559.2000.00895.x. - DOI - PubMed
-
- Hirabayashi S, Ohki K, Nakabayashi K, Ichikawa H, Momozawa Y, Okamura K, et al. ZNF384-related fusion genes define a subgroup of childhood B-cell precursor acute lymphoblastic leukemia with a characteristic immunotype. Haematologica. 2017;102:118–29. doi: 10.3324/haematol.2016.151035. - DOI - PMC - PubMed
-
- Ohki K, Kiyokawa N, Saito Y, Hirabayashi S, Nakabayashi K, Ichikawa H, et al. Clinical and molecular characteristics of MEF2D fusion-positive B-cell precursor acute lymphoblastic leukemia in childhood, including a novel translocation resulting in MEF2D-HNRNPH1 gene fusion. Haematologica. 2019;104:128–37. doi: 10.3324/haematol.2017.186320. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

