An ACE2 Triple Decoy that neutralizes SARS-CoV-2 shows enhanced affinity for virus variants
- PMID: 34140558
- PMCID: PMC8211782
- DOI: 10.1038/s41598-021-91809-9
An ACE2 Triple Decoy that neutralizes SARS-CoV-2 shows enhanced affinity for virus variants
Abstract
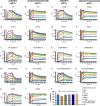
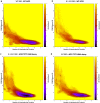
The SARS-CoV-2 variants replacing the first wave strain pose an increased threat by their potential ability to escape pre-existing humoral protection. An angiotensin converting enzyme 2 (ACE2) decoy that competes with endogenous ACE2 for binding of the SARS-CoV-2 spike receptor binding domain (S RBD) and inhibits infection may offer a therapeutic option with sustained efficacy against variants. Here, we used Molecular Dynamics (MD) simulation to predict ACE2 sequence substitutions that might increase its affinity for S RBD and screened candidate ACE2 decoys in vitro. The lead ACE2(T27Y/H34A)-IgG1FC fusion protein with enhanced S RBD affinity shows greater live SARS-CoV-2 virus neutralization capability than wild type ACE2. MD simulation was used to predict the effects of S RBD variant mutations on decoy affinity that was then confirmed by testing of an ACE2 Triple Decoy that included an additional enzyme activity-deactivating H374N substitution against mutated S RBD. The ACE2 Triple Decoy maintains high affinity for mutated S RBD, displays enhanced affinity for S RBD N501Y or L452R, and has the highest affinity for S RBD with both E484K and N501Y mutations, making it a viable therapeutic option for the prevention or treatment of SARS-CoV-2 infection with a high likelihood of efficacy against variants.
Conflict of interest statement
Authors with an ImmunityBio affiliation including Shiho Tanaka, Gard Nelson, C. Anders Olson, Oleksandr Buzko, Wendy Higashide, Annie Shin, Marcos Gonzalez, Patricia Spilman, Kayvan Niazi, Shahrooz Rabizadeh and Patrick Soon-Shiong designed, created or tested the ACE2 decoy, which may become a commercial product. Authors Justin Taft, Roosheel Patel, Sofija Buta, Ashley Richardson, and Dusan Bogunovic have no competing interests.
Figures


References
-
- Hodcroft, E.B., Domman, D.B., Snyder, D.J., Oguntuyo, Van Diest, M.V., Densmore, K.H., Schwalm, K.C., Femling, J., Carroll, J.L., Scott, R.S. et al. Emergence in late 2020 of multiple lineages of SARS-CoV-2 Spike protein variants affecting amino acid position 677. medRxiv 2021.02.12.21251658 (2021).
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

