Heterozygous OAS1 gain-of-function variants cause an autoinflammatory immunodeficiency
- PMID: 34145065
- PMCID: PMC8392508
- DOI: 10.1126/sciimmunol.abf9564
Heterozygous OAS1 gain-of-function variants cause an autoinflammatory immunodeficiency
Abstract
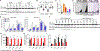
Analysis of autoinflammatory and immunodeficiency disorders elucidates human immunity and fosters the development of targeted therapies. Oligoadenylate synthetase 1 is a type I interferon-induced, intracellular double-stranded RNA (dsRNA) sensor that generates 2'-5'-oligoadenylate to activate ribonuclease L (RNase L) as a means of antiviral defense. We identified four de novo heterozygous OAS1 gain-of-function variants in six patients with a polymorphic autoinflammatory immunodeficiency characterized by recurrent fever, dermatitis, inflammatory bowel disease, pulmonary alveolar proteinosis, and hypogammaglobulinemia. To establish causality, we applied genetic, molecular dynamics simulation, biochemical, and cellular functional analyses in heterologous, autologous, and inducible pluripotent stem cell-derived macrophages and/or monocytes and B cells. We found that upon interferon-induced expression, OAS1 variant proteins displayed dsRNA-independent activity, which resulted in RNase L-mediated RNA cleavage, transcriptomic alteration, translational arrest, and dysfunction and apoptosis of monocytes, macrophages, and B cells. RNase L inhibition with curcumin modulated and allogeneic hematopoietic cell transplantation cured the disorder. Together, these data suggest that human OAS1 is a regulator of interferon-induced hyperinflammatory monocyte, macrophage, and B cell pathophysiology.
Copyright © 2021 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Competing interests:
We declare that none of the authors has competing financial or non-financial interests.
Figures




References
-
- Tangye SG, Al-Herz W, Bousfiha A, Chatila T, Cunningham-Rundles C, Etzioni A, Franco JL, Holland SM, Klein C, Morio T, Ochs HD, Oksenhendler E, Picard C, Puck J, Torgerson TR, Casanova JL, Sullivan KE, Human Inborn Errors of Immunity: 2019 Update on the Classification from the International Union of Immunological Societies Expert Committee. J Clin Immunol 40, 24–64 (2020). - PMC - PubMed
-
- Durandy A, Kracker S, Fischer A, Primary antibody deficiencies. Nat Rev Immunol 13, 519–533 (2013). - PubMed
-
- Liu Y, Jesus AA, Marrero B, Yang D, Ramsey SE, Sanchez GAM, Tenbrock K, Wittkowski H, Jones OY, Kuehn HS, Lee CR, DiMattia MA, Cowen EW, Gonzalez B, Palmer I, DiGiovanna JJ, Biancotto A, Kim H, Tsai WL, Trier AM, Huang Y, Stone DL, Hill S, Kim HJ, St Hilaire C, Gurprasad S, Plass N, Chapelle D, Horkayne-Szakaly I, Foell D, Barysenka A, Candotti F, Holland SM, Hughes JD, Mehmet H, Issekutz AC, Raffeld M, McElwee J, Fontana JR, Minniti CP, Moir S, Kastner DL, Gadina M, Steven AC, Wingfield PT, Brooks SR, Rosenzweig SD, Fleisher TA, Deng Z, Boehm M, Paller AS, Goldbach-Mansky R, Activated STING in a vascular and pulmonary syndrome. N Engl J Med 371, 507–518 (2014). - PMC - PubMed
-
- R. C. Group, Horby P, Lim WS, Emberson JR, Mafham M, Bell JL, Linsell L, Staplin N, Brightling C, Ustianowski A, Elmahi E, Prudon B, Green C, Felton T, Chadwick D, Rege K, Fegan C, Chappell LC, Faust SN, Jaki T, Jeffery K, Montgomery A, Rowan K, Juszczak E, Baillie JK, Haynes R, Landray MJ, Dexamethasone in Hospitalized Patients with Covid-19 - Preliminary Report. N Engl J Med, (2020). - PMC - PubMed
-
- Bronte V, Ugel S, Tinazzi E, Vella A, De Sanctis F, Cane S, Batani V, Trovato R, Fiore A, Petrova V, Hofer F, Barouni RM, Musiu C, Caligola S, Pinton L, Torroni L, Polati E, Donadello K, Friso S, Pizzolo F, Iezzi M, Facciotti F, Pelicci PG, Righetti D, Bazzoni P, Rampudda M, Comel AC, Mosaner W, Lunardi C, Olivieri O, Baricitinib restrains the immune dysregulation in severe COVID-19 patients. J Clin Invest, (2020). - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials