Indoleamine 2,3-dioxygenase 1 (IDO1): an up-to-date overview of an eclectic immunoregulatory enzyme
- PMID: 34145969
- PMCID: PMC9786828
- DOI: 10.1111/febs.16086
Indoleamine 2,3-dioxygenase 1 (IDO1): an up-to-date overview of an eclectic immunoregulatory enzyme
Abstract
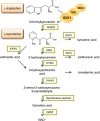
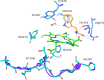
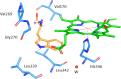
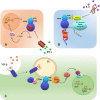
Indoleamine 2,3-dioxygenase 1 (IDO1) catalyzes the initial rate-limiting step in the degradation of the essential amino acid tryptophan along the kynurenine pathway. When discovered more than 50 years ago, IDO1 was thought to be an effector molecule capable of mediating a survival strategy based on the deprivation of bacteria and tumor cells of the essential amino acid tryptophan. Since 1998, when tryptophan catabolism was discovered to be crucially involved in the maintenance of maternal T-cell tolerance, IDO1 has become the focus of several laboratories around the world. Indeed, IDO1 is now considered as an authentic immune regulator not only in pregnancy, but also in autoimmune diseases, chronic inflammation, and tumor immunity. However, in the last years, a bulk of new information-including structural, biological, and functional evidence-on IDO1 has come to light. For instance, we now know that IDO1 has a peculiar conformational plasticity and, in addition to a complex and highly regulated catalytic activity, is capable of performing a nonenzymic function that reprograms the expression profile of immune cells toward a highly immunoregulatory phenotype. With this state-of-the-art review, we aimed at gathering the most recent information obtained for this eclectic protein as well as at highlighting the major unresolved questions.
Keywords: Src kinase; dendritic cells; early endosomes; indoleamine 2,3-dioxygenase 1; moonlighting protein; phosphatidylinositol 3-kinase; tryptophan metabolism.
© 2021 The Authors. The FEBS Journal published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Yamamoto S & Hayaishi O (1967) Tryptophan pyrrolase of rabbit intestine. D‐ and L‐tryptophan‐cleaving enzyme or enzymes. J Biol Chem 242, 5260–5266. - PubMed
-
- Munn DH, Zhou M, Attwood JT, Bondarev I, Conway SJ, Marshall B, Brown C & Mellor AL (1998) Prevention of allogeneic fetal rejection by tryptophan catabolism. Science 281, 1191–1193. - PubMed
-
- Thomas SR, Salahifar H, Mashima R, Hunt NH, Richardson DR & Stocker R (2001) Antioxidants inhibit indoleamine 2,3‐dioxygenase in IFN‐gamma‐activated human macrophages: posttranslational regulation by pyrrolidine dithiocarbamate. J Immunol 166, 6332–6340. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

