SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals
- PMID: 34155207
- PMCID: PMC8217552
- DOI: 10.1038/s41467-021-24007-w
SARS-CoV-2-mediated dysregulation of metabolism and autophagy uncovers host-targeting antivirals
Abstract
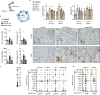
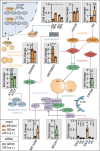
Viruses manipulate cellular metabolism and macromolecule recycling processes like autophagy. Dysregulated metabolism might lead to excessive inflammatory and autoimmune responses as observed in severe and long COVID-19 patients. Here we show that SARS-CoV-2 modulates cellular metabolism and reduces autophagy. Accordingly, compound-driven induction of autophagy limits SARS-CoV-2 propagation. In detail, SARS-CoV-2-infected cells show accumulation of key metabolites, activation of autophagy inhibitors (AKT1, SKP2) and reduction of proteins responsible for autophagy initiation (AMPK, TSC2, ULK1), membrane nucleation, and phagophore formation (BECN1, VPS34, ATG14), as well as autophagosome-lysosome fusion (BECN1, ATG14 oligomers). Consequently, phagophore-incorporated autophagy markers LC3B-II and P62 accumulate, which we confirm in a hamster model and lung samples of COVID-19 patients. Single-nucleus and single-cell sequencing of patient-derived lung and mucosal samples show differential transcriptional regulation of autophagy and immune genes depending on cell type, disease duration, and SARS-CoV-2 replication levels. Targeting of autophagic pathways by exogenous administration of the polyamines spermidine and spermine, the selective AKT1 inhibitor MK-2206, and the BECN1-stabilizing anthelmintic drug niclosamide inhibit SARS-CoV-2 propagation in vitro with IC50 values of 136.7, 7.67, 0.11, and 0.13 μM, respectively. Autophagy-inducing compounds reduce SARS-CoV-2 propagation in primary human lung cells and intestinal organoids emphasizing their potential as treatment options against COVID-19.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- WHO. https://www.who.int/emergencies/diseases/novel-coronavirus-2019. accessed 18 May 2021 (2021).
-
- Hoffmann, M. et al. Chloroquine does not inhibit infection of human lung cells with SARS-CoV-2. Nature, 10.1038/s41586-020-2575-3 (2020). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

