High-throughput Molecular Analysis of Pseudohypoparathyroidism 1b Patients Reveals Novel Genetic and Epigenetic Defects
- PMID: 34157100
- PMCID: PMC8677598
- DOI: 10.1210/clinem/dgab460
High-throughput Molecular Analysis of Pseudohypoparathyroidism 1b Patients Reveals Novel Genetic and Epigenetic Defects
Abstract
Context: Patients with pseudohypoparathyroidism type 1b (PHP1b) show disordered imprinting of the maternal GNAS allele or paternal uniparental disomy (UPD). Genetic deletions in STX16 or in upstream exons of GNAS are present in many familial but not sporadic cases.
Objective: Characterization of epigenetic and genetic defects in patients with PHP1b.
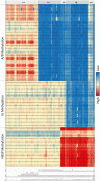
Design and patients: DNA from 84 subjects, including 26 subjects with sporadic PHP1b, 27 affected subjects and 17 unaffected and/or obligate gene carriers from 12 PHP1b families, 11 healthy individuals, and 3 subjects with PHP1a was subjected to quantitative pyrosequencing of GNAS differentially methylated regions (DMRs), microarray analysis, and microsatellite haplotype analysis.
Setting: Academic medical center.
Main outcome measurements: Molecular pathology of PHP1b.
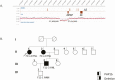
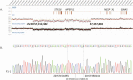
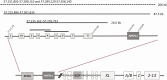
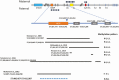
Results: Healthy subjects, unaffected family members and obligate carriers of paternal PHP1b alleles, and subjects with PHP1a showed normal methylation of all DMRs. All PHP1b subjects showed loss of methylation (LOM) at the exon A/B DMR. Affected members of 9 PHP1b kindreds showed LOM only at the exon A/B DMR, which was associated with a 3-kb deletion of STX16 exons 4 through 6 in 7 families and a novel deletion of STX16 and adjacent NEPEPL1 in 1 family. A novel NESP deletion was found in 1 of 2 other families with more extensive methylation defects. One sporadic PHP1b had UPD of 20q, 2 had 3-kb STX16 deletions, and 5 had apparent epigenetic mosaicism.
Conclusions: We found diverse patterns of defective methylation and identified novel or previously known mutations in 9 of 12 PHP1b families.
Keywords: GNAS; epigenetics; imprinting; parathyroid hormone; pseudohypoparathyroidism.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Endocrine Society. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures



Similar articles
-
Analysis of Multiple Families With Single Individuals Affected by Pseudohypoparathyroidism Type Ib (PHP1B) Reveals Only One Novel Maternally Inherited GNAS Deletion.J Bone Miner Res. 2016 Apr;31(4):796-805. doi: 10.1002/jbmr.2731. Epub 2015 Nov 14. J Bone Miner Res. 2016. PMID: 26479409 Free PMC article. Clinical Trial.
-
A Large Inversion Involving GNAS Exon A/B and All Exons Encoding Gsα Is Associated With Autosomal Dominant Pseudohypoparathyroidism Type Ib (PHP1B).J Bone Miner Res. 2017 Apr;32(4):776-783. doi: 10.1002/jbmr.3083. Epub 2017 Feb 24. J Bone Miner Res. 2017. PMID: 28084650 Free PMC article.
-
A Novel Familial PHP1B Variant With Incomplete Loss of Methylation at GNAS-A/B and Enhanced Methylation at GNAS-AS2.J Clin Endocrinol Metab. 2021 Aug 18;106(9):2779-2787. doi: 10.1210/clinem/dgab136. J Clin Endocrinol Metab. 2021. PMID: 33677588 Free PMC article.
-
Genetic and Epigenetic Characteristics of Autosomal Dominant Pseudohypoparathyroidism Type 1B: Case Reports and Literature Review.Horm Metab Res. 2021 Apr;53(4):225-235. doi: 10.1055/a-1341-9891. Epub 2021 Jan 29. Horm Metab Res. 2021. PMID: 33513624 Review.
-
Molecular Definition of Pseudohypoparathyroidism Variants.J Clin Endocrinol Metab. 2021 May 13;106(6):1541-1552. doi: 10.1210/clinem/dgab060. J Clin Endocrinol Metab. 2021. PMID: 33529330 Free PMC article. Review.
Cited by
-
GNAS locus: bone related diseases and mouse models.Front Endocrinol (Lausanne). 2023 Oct 18;14:1255864. doi: 10.3389/fendo.2023.1255864. eCollection 2023. Front Endocrinol (Lausanne). 2023. PMID: 37920253 Free PMC article. Review.
-
Novel PTH Gene Mutations Causing Isolated Hypoparathyroidism.J Clin Endocrinol Metab. 2022 May 17;107(6):e2449-e2458. doi: 10.1210/clinem/dgac086. J Clin Endocrinol Metab. 2022. PMID: 35165722 Free PMC article.
-
The long-range interaction between two GNAS imprinting control regions delineates pseudohypoparathyroidism type 1B pathogenesis.J Clin Invest. 2023 Apr 17;133(8):e167953. doi: 10.1172/JCI167953. J Clin Invest. 2023. PMID: 36853809 Free PMC article.
-
Bidirectional disruption of GNAS transcripts causes broad methylation defects in pseudohypoparathyroidism type 1B.Proc Natl Acad Sci U S A. 2025 Apr 22;122(16):e2423271122. doi: 10.1073/pnas.2423271122. Epub 2025 Apr 18. Proc Natl Acad Sci U S A. 2025. PMID: 40249781
-
Lack of GNAS Remethylation During Oogenesis May Be a Cause of Sporadic Pseudohypoparathyroidism Type Ib.J Clin Endocrinol Metab. 2022 Mar 24;107(4):e1610-e1619. doi: 10.1210/clinem/dgab830. J Clin Endocrinol Metab. 2022. PMID: 34791361 Free PMC article.
References
-
- Thiele S, Mantovani G, Barlier A, et al. From pseudohypoparathyroidism to inactivating PTH/PTHrP signalling disorder (iPPSD), a novel classification proposed by the EuroPHP network. Eur J Endocrinol. 2016;175(6):P1-P17. - PubMed
-
- Farfel Z, Brickman AS, Kaslow HR, Brothers VM, Bourne HR. Defect of receptor-cyclase coupling protein in pseudohypoparathyroidism. N.Engl.J.Med. 1980;303:237-242. - PubMed
-
- Levine MA, Downs RW Jr, Singer M, Marx SJ, Aurbach GD, Spiegel AM. Deficient activity of guanine nucleotide regulatory protein in erythrocytes from patients with pseudohypoparathyroidism. Biochem Biophys Res Commun. 1980;94(4):1319-1324. - PubMed

