Identification of presented SARS-CoV-2 HLA class I and HLA class II peptides using HLA peptidomics
- PMID: 34166618
- PMCID: PMC8185308
- DOI: 10.1016/j.celrep.2021.109305
Identification of presented SARS-CoV-2 HLA class I and HLA class II peptides using HLA peptidomics
Abstract
The human leukocyte antigen (HLA)-bound viral antigens serve as an immunological signature that can be selectively recognized by T cells. As viruses evolve by acquiring mutations, it is essential to identify a range of presented viral antigens. Using HLA peptidomics, we are able to identify severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)-derived peptides presented by highly prevalent HLA class I (HLA-I) molecules by using infected cells as well as overexpression of SARS-CoV-2 genes. We find 26 HLA-I peptides and 36 HLA class II (HLA-II) peptides. Among the identified peptides, some are shared between different cells and some are derived from out-of-frame open reading frames (ORFs). Seven of these peptides were previously shown to be immunogenic, and we identify two additional immunoreactive peptides by using HLA multimer staining. These results may aid the development of the next generation of SARS-CoV-2 vaccines based on presented viral-specific antigens that span several of the viral genes.
Keywords: HLA; Peptides; Peptidomics; SARS-CoV-2; immuno-reactive; out-of-frame-ORFs.
Copyright © 2021 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests Patent applications have been filed on SARS-CoV-2 peptides (Y.S., A.N., and S.K.). The remaining authors declare no competing interests.
Figures


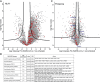



References
-
- Abelin J.G., Harjanto D., Malloy M., Suri P., Colson T., Goulding S.P., Creech A.L., Serrano L.R., Nasir G., Nasrullah Y. Defining HLA-II Ligand Processing and Binding Rules with Mass Spectrometry Enhances Cancer Epitope Prediction. Immunity. 2019;51:766–779.e17. - PubMed
-
- Andreatta M., Lund O., Nielsen M. Simultaneous alignment and clustering of peptide data using a Gibbs sampling approach. Bioinformatics. 2013;29:8–14. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

