Intratumor CMS Heterogeneity Impacts Patient Prognosis in Localized Colon Cancer
- PMID: 34168047
- PMCID: PMC8974433
- DOI: 10.1158/1078-0432.CCR-21-0529
Intratumor CMS Heterogeneity Impacts Patient Prognosis in Localized Colon Cancer
Abstract
Purpose: The consensus molecular subtypes (CMS) represent a significant advance in the understanding of intertumor heterogeneity in colon cancer. Intratumor heterogeneity (ITH) is the new frontier for refining prognostication and understanding treatment resistance. This study aims at deciphering the transcriptomic ITH of colon cancer and understanding its potential prognostic implications.
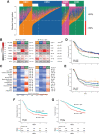
Experimental design: We deconvoluted the transcriptomic profiles of 1,779 tumors from the PETACC8 trial and 155 colon cancer cell lines as weighted sums of the four CMSs, using the Weighted In Silico Pathology (WISP) algorithm. We assigned to each tumor and cell line a combination of up to three CMS subtypes with a threshold above 20%.
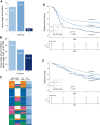
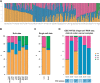
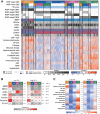
Results: Over 55% of tumors corresponded to mixtures of at least two CMSs, demonstrating pervasive ITH in colon cancer. Of note, ITH was associated with shorter disease-free survival (DFS) and overall survival, [HR, 1.34; 95% confidence interval (CI; 1.12-1.59), 1.40, 95% CI (1.14-1.71), respectively]. Moreover, we uncovered specific combinations of CMS associated with dismal prognosis. In multivariate analysis, ITH represents the third parameter explaining DFS variance, after T and N stages. At a cellular level, combined WISP and single-cell transcriptomic analysis revealed that most colon cancer cell lines are a mixture of cells falling into different CMSs, indicating that ITH may correspond to distinct functional statuses of colon cancer cells.
Conclusions: This study shows that CMS-based transcriptomic ITH is frequent in colon cancer and impacts its prognosis. CMS-based transcriptomic ITH may correspond to distinct functional statuses of colon cancer cells, suggesting plasticity between CMS-related cell populations. Transcriptomic ITH deserves further assessment in the context of personalized medicine.
©2021 The Authors; Published by the American Association for Cancer Research.
Conflict of interest statement
C. Lepage reports personal fees from Bayer, Amgen, Ipsen, Pierre Fabre, and Novartis outside the submitted work. V. Boige reports grants, personal fees, and non-financial support from Merck Serono; personal fees and non-financial support from Bayer, Roche, Ipsen, Merck MSD, and Amgen; non-financial support from Sanofi; and personal fees from BMS, Eisai, and Novartis outside the submitted work. J.-F. Emile reports personal fees from Pierre Fabre, Merck Sharp & Dohme, Merck Serono, Amgen, Novartis, and Bristol Myers Squibb outside the submitted work. P. Laurent-Puig reports grants from Federeation francophones de cancerologie disgestive and Ligue National de luttte contre le cancer during the conduct of the study, as well as personal fees from Amgen, Boehringer Ingelheim, Sanofi, MSD, Imedex, Pierre Fabre, and Institut Servier; grants and personal fees from Biocartis, BMS, and Roche; and grants from Institut Roche, Bio-Rad, and Servier outside the submitted work. No disclosures were reported by the other authors.
Figures
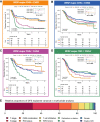

References
-
- Allemani C, Matsuda T, Di Carlo V, Harewood R, Matz M, Nikšić M, et al. Global surveillance of trends in cancer survival 2000–14 (CONCORD-3): analysis of individual records for 37 513 025 patients diagnosed with one of 18 cancers from 322 population-based registries in 71 countries. Lancet 2018;391:1023–75. - PMC - PubMed
-
- Van Cutsem E, Cervantes A, Adam R, Sobrero A, Van Krieken JH, Aderka D, et al. ESMO consensus guidelines for the management of patients with metastatic colorectal cancer. Ann Oncol 2016;27:1386–422. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

