Highly-potent, synthetic APOBEC3s restrict HIV-1 through deamination-independent mechanisms
- PMID: 34170969
- PMCID: PMC8266076
- DOI: 10.1371/journal.ppat.1009523
Highly-potent, synthetic APOBEC3s restrict HIV-1 through deamination-independent mechanisms
Abstract
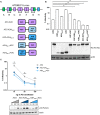
The APOBEC3 (A3) genes encode cytidine deaminase proteins with potent antiviral and anti-retroelement activity. This locus is characterized by duplication, recombination, and deletion events that gave rise to the seven A3s found in primates. These include three single deaminase domain A3s (A3A, A3C, and A3H) and four double deaminase domain A3s (A3B, A3D, A3F, and A3G). The most potent of the A3 proteins against HIV-1 is A3G. However, it is not clear if double deaminase domain A3s have a generalized functional advantage to restrict HIV-1. In order to test whether superior restriction factors could be created by genetically linking single A3 domains into synthetic double domains, we linked A3C and A3H single domains in novel combinations. We found that A3C/A3H double domains acquired enhanced antiviral activity that is at least as potent, if not better than, A3G. Although these synthetic double domain A3s package into budding virions more efficiently than their respective single domains, this does not fully explain their gain of antiviral potency. The antiviral activity is conferred both by cytidine-deaminase dependent and independent mechanisms, with the latter correlating to an increase in RNA binding affinity. T cell lines expressing this A3C-A3H super restriction factor are able to control replicating HIV-1ΔVif infection to similar levels as A3G. Together, these data show that novel combinations of A3 domains are capable of gaining potent antiviral activity to levels similar to the most potent genome-encoded A3s, via a primarily non-catalytic mechanism.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
APOBEC3C Tandem Domain Proteins Create Super Restriction Factors against HIV-1.mBio. 2020 Apr 28;11(2):e00737-20. doi: 10.1128/mBio.00737-20. mBio. 2020. PMID: 32345636 Free PMC article.
-
Sequence and structural determinants of human APOBEC3H deaminase and anti-HIV-1 activities.Retrovirology. 2015 Jan 22;12:3. doi: 10.1186/s12977-014-0130-8. Retrovirology. 2015. PMID: 25614027 Free PMC article.
-
Structural basis of substrate specificity in human cytidine deaminase family APOBEC3s.J Biol Chem. 2021 Aug;297(2):100909. doi: 10.1016/j.jbc.2021.100909. Epub 2021 Jun 24. J Biol Chem. 2021. PMID: 34171358 Free PMC article.
-
Role of the single deaminase domain APOBEC3A in virus restriction, retrotransposition, DNA damage and cancer.J Gen Virol. 2016 Jan;97(1):1-17. doi: 10.1099/jgv.0.000320. Epub 2015 Oct 20. J Gen Virol. 2016. PMID: 26489798 Free PMC article. Review.
-
Structural Insights into APOBEC3-Mediated Lentiviral Restriction.Viruses. 2020 May 27;12(6):587. doi: 10.3390/v12060587. Viruses. 2020. PMID: 32471198 Free PMC article. Review.
Cited by
-
Stability of APOBEC3F in the Presence of the APOBEC3 Antagonist HIV-1 Vif Increases at the Expense of Co-Expressed APOBEC3H Haplotype I.Viruses. 2023 Feb 7;15(2):463. doi: 10.3390/v15020463. Viruses. 2023. PMID: 36851677 Free PMC article.
-
Revisiting the MMTV Zoonotic Hypothesis to Account for Geographic Variation in Breast Cancer Incidence.Viruses. 2022 Mar 9;14(3):559. doi: 10.3390/v14030559. Viruses. 2022. PMID: 35336966 Free PMC article.
-
Mechanistic Interplay between HIV-1 Reverse Transcriptase Enzyme Kinetics and Host SAMHD1 Protein: Viral Myeloid-Cell Tropism and Genomic Mutagenesis.Viruses. 2022 Jul 26;14(8):1622. doi: 10.3390/v14081622. Viruses. 2022. PMID: 35893688 Free PMC article. Review.
-
Low prevalence of HIV in the northern Cameroon: contribution of some AIDS restriction genes and potential implications for gene therapy.Front Genet. 2024 Sep 13;15:1447971. doi: 10.3389/fgene.2024.1447971. eCollection 2024. Front Genet. 2024. PMID: 39346778 Free PMC article.
-
Molecular mechanism for regulating APOBEC3G DNA editing function by the non-catalytic domain.Nat Commun. 2024 Oct 10;15(1):8773. doi: 10.1038/s41467-024-52671-1. Nat Commun. 2024. PMID: 39389938 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

