Harnessing clinical annotations to improve deep learning performance in prostate segmentation
- PMID: 34170972
- PMCID: PMC8232529
- DOI: 10.1371/journal.pone.0253829
Harnessing clinical annotations to improve deep learning performance in prostate segmentation
Abstract
Purpose: Developing large-scale datasets with research-quality annotations is challenging due to the high cost of refining clinically generated markup into high precision annotations. We evaluated the direct use of a large dataset with only clinically generated annotations in development of high-performance segmentation models for small research-quality challenge datasets.
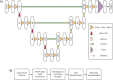
Materials and methods: We used a large retrospective dataset from our institution comprised of 1,620 clinically generated segmentations, and two challenge datasets (PROMISE12: 50 patients, ProstateX-2: 99 patients). We trained a 3D U-Net convolutional neural network (CNN) segmentation model using our entire dataset, and used that model as a template to train models on the challenge datasets. We also trained versions of the template model using ablated proportions of our dataset, and evaluated the relative benefit of those templates for the final models. Finally, we trained a version of the template model using an out-of-domain brain cancer dataset, and evaluated the relevant benefit of that template for the final models. We used five-fold cross-validation (CV) for all training and evaluation across our entire dataset.
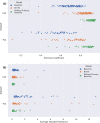
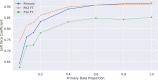
Results: Our model achieves state-of-the-art performance on our large dataset (mean overall Dice 0.916, average Hausdorff distance 0.135 across CV folds). Using this model as a pre-trained template for refining on two external datasets significantly enhanced performance (30% and 49% enhancement in Dice scores respectively). Mean overall Dice and mean average Hausdorff distance were 0.912 and 0.15 for the ProstateX-2 dataset, and 0.852 and 0.581 for the PROMISE12 dataset. Using even small quantities of data to train the template enhanced performance, with significant improvements using 5% or more of the data.
Conclusion: We trained a state-of-the-art model using unrefined clinical prostate annotations and found that its use as a template model significantly improved performance in other prostate segmentation tasks, even when trained with only 5% of the original dataset.
Conflict of interest statement
LSM and AMP report a financial interest in Avenda Health outside the submitted work. BT reports IP-related royalties from Philips outside the submitted work. The NIH has cooperative research and development agreements with NVIDIA, Philips, Siemens, Xact Robotics, Celsion Corp, and Boston Scientific outside the submitted work. The NIH has research partnerships with Angiodynamics, ArciTrax, and Exact Imaging outside the submitted work. CWA has received research equipment from NVIDIA Corporation, outside the submitted work. No commercial funding or equipment was used in the execution of this study. No other authors have competing interests to disclose.
Figures

References
-
- Invivo-Philips. DynaCAD Prostate Advanced visualization for prostate MRI analysis | Philips Healthcare. [cited 26 Apr 2021]. Available: https://www.usa.philips.com/healthcare/product/HC784029/dynacad-prostate
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

