New insights into the intraspecific cytoplasmic DNA diversity, maternal lineages classification and conservation issues of Tunisian pearl millet landraces
- PMID: 34177320
- PMCID: PMC8215465
- DOI: 10.5511/plantbiotechnology.20.0831a
New insights into the intraspecific cytoplasmic DNA diversity, maternal lineages classification and conservation issues of Tunisian pearl millet landraces
Abstract
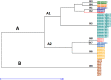
Tunisian pearl millet (Pennisetum glaucum L.) landraces are still growing in contrasting agro-ecological environments and are considered potentially useful for national and international breeders. Despite its genetic potential, the cropping areas of this species are still limited and scattered which increases the risk of genetic erosion. The chloroplast DNA polymorphism and maternal lineages classification of forty nine pearl millet landraces representing seven populations covering the main distribution area of this crop in Tunisia were undertaken based on informative cpSSR molecular markers. A total of 21 alleles combining to 9 haplotypes were detected with a mean value of 3.5 alleles per locus and a haplotype genetic diversity (Hd) of 0.82. The number of chloroplast haplotypes per population ranged from 1 to 4 with an average of 1.28. The haplotypes median-joining network and UPGMA analyses revealed two probable ancestral maternal lineages with a differential pearl millet seed-exchange rate between the investigated areas. Northern and Central populations presented unique genetic backgrounds while historical farmers' practices in the South-East area resulted in the isolation of their own local landraces. The genetic evidences strongly support at least two introduction origins of pearl millet in Tunisia, one in the North and the other in the South followed by distinct local dispersal histories. Complementary in-situ and ex-situ conservation strategies taking into account the conservation of the maternal lineage cytoplasmic diversity are required. The investigated chloroplast SSRs provide useful molecular markers which could be used in further genetic studies and breeding surveys of pearl millet genetic resources.
Keywords: Pennisetum glaucum L; conservation; cpSSR; cytoplasmic DNA; maternal lineages.
© 2021 Japanese Society for Plant Biotechnology.
Figures

References
-
- Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16: 37–48 - PubMed
-
- Ben Romdhane M, Riahi L, Selmi A, Jardak R, Bouajila A, Ghorbel A, Zoghlami N (2017) Low genetic differentiation and evidence of gene flow among barley landrace populations in Tunisia. Crop Sci 57: 1–9
-
- Dias-Martins AM, Pessanha KLF, Pacheco S, Rodrigues JAS, Carvalho CWP (2018) Potential use of pearl millet (Pennisetum glaucum (L.) R. Br.) in Brazil: Food security, processing, health benefits and nutritional products. Food Res Int 109: 175–186 - PubMed
-
- Eliades NG, Eliades DG (2009) Haplotype Analysis: Software for Analysis of Haplotypes Data. Distributed by the Authors. Forest Genetics and Forest Tree Breeding, Georg-Augst University Goettingen, Germany
LinkOut - more resources
Full Text Sources
