Topological data analysis distinguishes parameter regimes in the Anderson-Chaplain model of angiogenesis
- PMID: 34181657
- PMCID: PMC8270459
- DOI: 10.1371/journal.pcbi.1009094
Topological data analysis distinguishes parameter regimes in the Anderson-Chaplain model of angiogenesis
Abstract
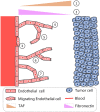
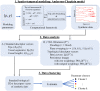
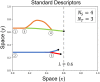
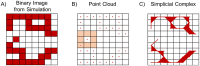
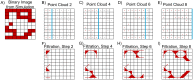
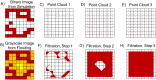
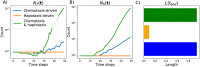
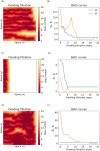
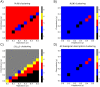
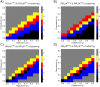
Angiogenesis is the process by which blood vessels form from pre-existing vessels. It plays a key role in many biological processes, including embryonic development and wound healing, and contributes to many diseases including cancer and rheumatoid arthritis. The structure of the resulting vessel networks determines their ability to deliver nutrients and remove waste products from biological tissues. Here we simulate the Anderson-Chaplain model of angiogenesis at different parameter values and quantify the vessel architectures of the resulting synthetic data. Specifically, we propose a topological data analysis (TDA) pipeline for systematic analysis of the model. TDA is a vibrant and relatively new field of computational mathematics for studying the shape of data. We compute topological and standard descriptors of model simulations generated by different parameter values. We show that TDA of model simulation data stratifies parameter space into regions with similar vessel morphology. The methodologies proposed here are widely applicable to other synthetic and experimental data including wound healing, development, and plant biology.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

