Subtractive proteomics approach to Unravel the druggable proteins of the emerging pathogen Waddlia chondrophila and drug repositioning on its MurB protein
- PMID: 34195427
- PMCID: PMC8239728
- DOI: 10.1016/j.heliyon.2021.e07320
Subtractive proteomics approach to Unravel the druggable proteins of the emerging pathogen Waddlia chondrophila and drug repositioning on its MurB protein
Abstract
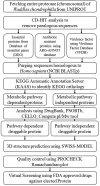
Waddlia chondrophila is an emerging pathogen that has been implicated in numerous unpropitious pregnancy events in humans and ruminants. Taking into account its association with abortigenic events, possible modes of transmission, and future risk, immediate clinical measures are required to prevent widespread damage caused by this organism and hence this study. Here, a subtractive proteomics approach was employed to identify druggable proteins of W. chondrophila. Considering the essential genes, antibiotic resistance proteins, and virulence factors, 676 unique important proteins were initially identified for this bacterium. Afterward, NCBI BLASTp performed against human proteome identified 223 proteins that were further pushed into KEGG Automatic Annotation Server (KAAS) for automatic annotation. Using the information from the Kyoto Encyclopedia of Genes and Genomes (KEGG) database 14 Waddlia specific metabolic pathways were identified with respect to humans. Analyzing the data from KAAS and KEGG databases, forty-eight metabolic pathway-dependent, and seventy metabolic pathway independent proteins were identified. Standalone BLAST search against DrugBank FDA approved drug targets revealed eight proteins that are finally considered druggable proteins. Prediction of three-dimensional structures was done for the eight proteins through homology modeling and the Ramachandran plot model showed six models as a valid prediction. Finally, virtual screening against MurB protein was performed using FDA approved drugs to employ the drug repositioning strategy. Three drugs showed promising docking results that can be used for therapeutic purposes against W. chondrophila following the clinical validation of the study.
Keywords: DrugBank; KEGG; Modelling; Molecular docking; Subtractive proteomics; Waddlia chondrophila.
© 2021 The Authors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Pharmacophore modelling based virtual screening and molecular dynamics identified the novel inhibitors and drug targets against Waddlia chondrophila.Sci Rep. 2024 Jun 12;14(1):13472. doi: 10.1038/s41598-024-63555-1. Sci Rep. 2024. PMID: 38866811 Free PMC article.
-
In Silico Subtractive Proteomics and Molecular Docking Approaches for the Identification of Novel Inhibitors against Streptococcus pneumoniae Strain D39.Life (Basel). 2023 May 4;13(5):1128. doi: 10.3390/life13051128. Life (Basel). 2023. PMID: 37240772 Free PMC article.
-
Identification of immunogenic proteins of Waddlia chondrophila.PLoS One. 2012;7(1):e28605. doi: 10.1371/journal.pone.0028605. Epub 2012 Jan 4. PLoS One. 2012. PMID: 22238579 Free PMC article.
-
Undressing of Waddlia chondrophila to enrich its outer membrane proteins to develop a new species-specific ELISA.New Microbes New Infect. 2014 Jan;2(1):13-24. doi: 10.1002/2052-2975.26. Epub 2014 Jan 16. New Microbes New Infect. 2014. PMID: 25356333 Free PMC article.
-
Waddlia chondrophila: from biology to pathogenicity.Microbes Infect. 2013 Dec;15(14-15):1033-41. doi: 10.1016/j.micinf.2013.09.010. Epub 2013 Oct 18. Microbes Infect. 2013. PMID: 24141085 Review.
Cited by
-
Absorption wavelength (TD-DFT) and adsorption of metal chalcogen clusters with methyl nicotinate: Structural, electronic, IRI, SERS, pharmacological and antiviral studies (HIV and omicron).Heliyon. 2023 May 13;9(5):e16066. doi: 10.1016/j.heliyon.2023.e16066. eCollection 2023 May. Heliyon. 2023. PMID: 37234664 Free PMC article.
-
Structural, electronic, intermolecular interaction, reactivity, vibrational spectroscopy, charge transfer, Hirshfeld surface analysis, pharmacological and hydropathy plot on 5-Bromo nicotinic acid - Antiviral study (Hepatitis A, B, and C).Heliyon. 2023 Sep 7;9(9):e19965. doi: 10.1016/j.heliyon.2023.e19965. eCollection 2023 Sep. Heliyon. 2023. PMID: 37809934 Free PMC article.
-
Proteome-wide identification of druggable targets and inhibitors for multidrug-resistant Pseudomonas aeruginosa using an integrative subtractive proteomics and virtual screening approach.Heliyon. 2025 Feb 10;11(4):e42584. doi: 10.1016/j.heliyon.2025.e42584. eCollection 2025 Feb 28. Heliyon. 2025. PMID: 40066032 Free PMC article.
References
-
- Woolhouse M.E.J. Population biology of emerging and re-emerging pathogens. Trends Microbiol. 2002;10 - PubMed
-
- McMichael A.J., Woodruff R.E., Hales S. Climate change and human health: present and future risks. Lancet. 2006;367:859–869. - PubMed
-
- Ligon B.L. Infectious diseases that pose specific challenges after natural disasters: a review. Semin. Pediatr. Infect. Dis. 2006;17:36–45. - PubMed
-
- Millar B.C., Moore J.E. Emerging pathogens in infectious diseases: definitions, causes and trends. Rev. Med. Microbiol. 2006;17:101–106.
LinkOut - more resources
Full Text Sources
Research Materials

