From Residues to Added-Value Bacterial Biopolymers as Nanomaterials for Biomedical Applications
- PMID: 34200068
- PMCID: PMC8228158
- DOI: 10.3390/nano11061492
From Residues to Added-Value Bacterial Biopolymers as Nanomaterials for Biomedical Applications
Abstract
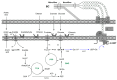
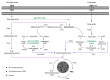
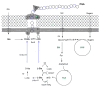
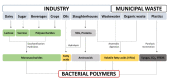
Bacterial biopolymers are naturally occurring materials comprising a wide range of molecules with diverse chemical structures that can be produced from renewable sources following the principles of the circular economy. Over the last decades, they have gained substantial interest in the biomedical field as drug nanocarriers, implantable material coatings, and tissue-regeneration scaffolds or membranes due to their inherent biocompatibility, biodegradability into nonhazardous disintegration products, and their mechanical properties, which are similar to those of human tissues. The present review focuses upon three technologically advanced bacterial biopolymers, namely, bacterial cellulose (BC), polyhydroxyalkanoates (PHA), and γ-polyglutamic acid (PGA), as models of different carbon-backbone structures (polysaccharides, polyesters, and polyamides) produced by bacteria that are suitable for biomedical applications in nanoscale systems. This selection models evidence of the wide versatility of microorganisms to generate biopolymers by diverse metabolic strategies. We highlight the suitability for applied sustainable bioprocesses for the production of BC, PHA, and PGA based on renewable carbon sources and the singularity of each process driven by bacterial machinery. The inherent properties of each polymer can be fine-tuned by means of chemical and biotechnological approaches, such as metabolic engineering and peptide functionalization, to further expand their structural diversity and their applicability as nanomaterials in biomedicine.
Keywords: bacterial cellulose; bacterial polymers; biomedical applications; biopolymer functionalization; polyhydroxyalkanoates; upcycled polymers; γ-polyglutamic acid.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

