Detection of Genomic Regions with Pleiotropic Effects for Growth and Carcass Quality Traits in the Rubia Gallega Cattle Breed
- PMID: 34200089
- PMCID: PMC8227173
- DOI: 10.3390/ani11061682
Detection of Genomic Regions with Pleiotropic Effects for Growth and Carcass Quality Traits in the Rubia Gallega Cattle Breed
Abstract
The breeding scheme in the Rubia Gallega cattle population is based upon traits measured in farms and slaughterhouses. In recent years, genomic evaluation has been implemented by using a ssGBLUP (single-step Genomic Best Linear Unbiased Prediction). This procedure can reparameterized to perform ssGWAS (single-step Genome Wide Association Studies) by backsolving the SNP (single nucleotide polymorphisms) effects. Therefore, the objective of this study was to identify genomic regions associated with the genetic variability in growth and carcass quality traits. We implemented a ssGBLUP by using a database that included records for Birth Weight (BW-327,350 records-), Weaning Weight (WW-83,818-), Cold Carcass Weight (CCW-91,621-), Fatness (FAT-91,475-) and Conformation (CON-91,609-). The pedigree included 464,373 individuals, 2449 of which were genotyped. After a process of filtering, we ended up using 43,211 SNP markers. We used the GBLUP and SNPBLUP model equivalences to obtain the effects of the SNPs and then calculated the percentage of variance explained by the regions of the genome between 1 Mb. We identified 7 regions of the genome for CCW; 8 regions for BW, WW, FAT and 9 regions for CON, which explained the percentage of variance above 0.5%. Furthermore, a number of the genome regions had pleiotropic effects, located at: BTA1 (131-132 Mb), BTA2 (1-11 Mb), BTA3 (32-33 Mb), BTA6 (36-38 Mb), BTA16 (24-26 Mb), and BTA 21 (56-57 Mb). These regions contain, amongst others, the following candidate genes: NCK1, MSTN, KCNA3, LCORL, NCAPG, and RIN3.
Keywords: GWAS; SNP; beef cattle; candidate genes; pleiotropy; single-step GBLUP.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
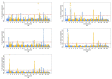
Similar articles
-
Reaffirmation of known major genes and the identification of novel candidate genes associated with carcass-related metrics based on whole genome sequence within a large multi-breed cattle population.BMC Genomics. 2019 Sep 18;20(1):720. doi: 10.1186/s12864-019-6071-9. BMC Genomics. 2019. PMID: 31533623 Free PMC article.
-
Genetic Architecture of Carcass and Meat Quality Traits in Montana Tropical® Composite Beef Cattle.Front Genet. 2020 Feb 27;11:123. doi: 10.3389/fgene.2020.00123. eCollection 2020. Front Genet. 2020. PMID: 32180796 Free PMC article.
-
Genome-Wide Association Study between Single Nucleotide Polymorphisms and Flight Speed in Nellore Cattle.PLoS One. 2016 Jun 14;11(6):e0156956. doi: 10.1371/journal.pone.0156956. eCollection 2016. PLoS One. 2016. PMID: 27300296 Free PMC article.
-
Comparison of conventional BLUP and single-step genomic BLUP evaluations for yearling weight and carcass traits in Hanwoo beef cattle using single trait and multi-trait models.PLoS One. 2019 Oct 14;14(10):e0223352. doi: 10.1371/journal.pone.0223352. eCollection 2019. PLoS One. 2019. PMID: 31609979 Free PMC article.
-
Genome-wide association and genotype by environment interactions for growth traits in U.S. Gelbvieh cattle.BMC Genomics. 2019 Dec 4;20(1):926. doi: 10.1186/s12864-019-6231-y. BMC Genomics. 2019. PMID: 31801456 Free PMC article.
Cited by
-
Selective Sweeps in Cattle Genomes in Response to the Influence of Urbanization and Environmental Contamination.Genes (Basel). 2023 Nov 15;14(11):2083. doi: 10.3390/genes14112083. Genes (Basel). 2023. PMID: 38003026 Free PMC article.
-
Genome-wide association studies and genetic architecture of carcass traits in Angus beef cattle using imputed whole-genome sequences data.Genet Sel Evol. 2025 Jun 1;57(1):26. doi: 10.1186/s12711-025-00970-6. Genet Sel Evol. 2025. PMID: 40452020 Free PMC article.
-
Multiple Trait Bayesian Analysis of Partitioned Genetic Trends Accounting for Uncertainty in Genetic Parameters. An Example With the Pirenaica and Rubia Gallega Beef Cattle Breeds.J Anim Breed Genet. 2025 Jul;142(4):454-462. doi: 10.1111/jbg.12918. Epub 2024 Dec 19. J Anim Breed Genet. 2025. PMID: 39698940 Free PMC article.
References
-
- Gobierno de España, Ministerio de Agricultura Pesca y Alimentacion Raza Bovina RUBIA GALLEGA. [(accessed on 1 April 2021)]; Available online: https://www.mapa.gob.es/es/ganaderia/temas/zootecnia/razas-ganaderas/raz....
-
- Cantalapiedra J., Rodríguez M., Payán R., Camiña M., Iglesias A. Evolución Morfológica de la Raza Rubia Gallega Basada en Sus Medidas Zoométricas e Índices Etnológicos. In: Marta-Costa A.A., Tibério M.L., Payan-Carreira R., editors. Raças Autóctones no Espaço Ibérico: Um Recurso Sustentável. Universidade de Trás-os-Montes e Alto Douro; Vila Real, Spain: 2016. [(accessed on 15 March 2021)]. pp. 65–69. Available online: https://www.ruralbit.com/client_manager/files/1448372206-1261.pdf.
-
- Legarra A., Christensen O.F., Aguilar I., Misztal I. Single Step, a general approach for genomic selection. Livest. Sci. 2014;166:54–65. doi: 10.1016/j.livsci.2014.04.029. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

