ACE2-Independent Interaction of SARS-CoV-2 Spike Protein with Human Epithelial Cells Is Inhibited by Unfractionated Heparin
- PMID: 34200372
- PMCID: PMC8229176
- DOI: 10.3390/cells10061419
ACE2-Independent Interaction of SARS-CoV-2 Spike Protein with Human Epithelial Cells Is Inhibited by Unfractionated Heparin
Abstract
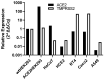
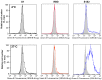
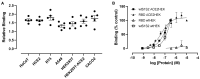
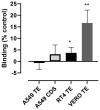
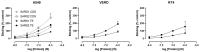
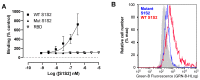
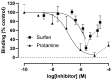
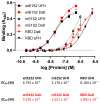
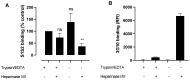
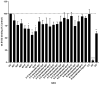
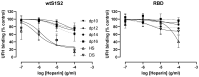
Coronaviruses such as SARS-CoV-2, which is responsible for COVID-19, depend on virus spike protein binding to host cell receptors to cause infection. The SARS-CoV-2 spike protein binds primarily to ACE2 on target cells and is then processed by membrane proteases, including TMPRSS2, leading to viral internalisation or fusion with the plasma membrane. It has been suggested, however, that receptors other than ACE2 may be involved in virus binding. We have investigated the interactions of recombinant versions of the spike protein with human epithelial cell lines that express low/very low levels of ACE2 and TMPRSS2 in a proxy assay for interaction with host cells. A tagged form of the spike protein containing the S1 and S2 regions bound in a temperature-dependent manner to all cell lines, whereas the S1 region alone and the receptor-binding domain (RBD) interacted only weakly. Spike protein associated with cells independently of ACE2 and TMPRSS2, while RBD required the presence of high levels of ACE2 for interaction. As the spike protein has previously been shown to bind heparin, a soluble glycosaminoglycan, we tested the effects of various heparins on ACE2-independent spike protein interaction with cells. Unfractionated heparin inhibited spike protein interaction with an IC50 value of <0.05 U/mL, whereas two low-molecular-weight heparins were less effective. A mutant form of the spike protein, lacking the arginine-rich putative furin cleavage site, interacted only weakly with cells and had a lower affinity for unfractionated and low-molecular-weight heparin than the wild-type spike protein. This suggests that the furin cleavage site might also be a heparin-binding site and potentially important for interactions with host cells. The glycosaminoglycans heparan sulphate and dermatan sulphate, but not chondroitin sulphate, also inhibited the binding of spike protein, indicating that it might bind to one or both of these glycosaminoglycans on the surface of target cells.
Keywords: SARS-CoV-2; glycosaminoglycan; heparin; spike protein.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures

References
-
- Hoffmann M., Kleine-Weber H., Schroeder S., Krüger N., Herrler T., Erichsen S., Schiergens T.S., Herrler G., Wu N.-H., Nitsche A., et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell. 2020;181:271–280. doi: 10.1016/j.cell.2020.02.052. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

