Anti-SARS-CoV-2 Vaccines and Monoclonal Antibodies Facing Viral Variants
- PMID: 34207378
- PMCID: PMC8234553
- DOI: 10.3390/v13061171
Anti-SARS-CoV-2 Vaccines and Monoclonal Antibodies Facing Viral Variants
Abstract
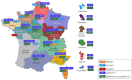
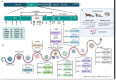
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is genetically variable, allowing it to adapt to various hosts including humans. Indeed, SARS-CoV-2 has accumulated around two mutations per genome each month. The first relevant event in this context was the occurrence of the mutant D614G in the Spike gene. Moreover, several variants have emerged, including the well-characterized 20I/501Y.V1, 20H/501Y.V2, and 20J/501Y.V3 strains, in addition to those that have been detected within clusters, such as 19B/501Y or 20C/655Y in France. Mutants have also emerged in animals, including a variant transmitted to humans, namely, the Mink variant detected in Denmark. The emergence of these variants has affected the transmissibility of the virus (for example, 20I/501Y.V1, which was up to 82% more transmissible than other preexisting variants), its severity, and its ability to escape natural, adaptive, vaccine, and therapeutic immunity. In this respect, we review the literature on variants that have currently emerged, and their effect on vaccines and therapies, and, in particular, monoclonal antibodies (mAbs). The emergence of SARS-CoV-2 variants must be examined to allow effective preventive and curative control strategies to be developed.
Keywords: SARS-CoV-2; antibody; emergence; immunity; mutations; vaccine; variant.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
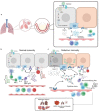
- Hoffmann M., Kleine-Weber H., Schroeder S., Krüger N., Herrler T., Erichsen S., Schiergens T.S., Herrler G., Wu N.-H., Nitsche A., et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell. 2020;181:271–280.e8. doi: 10.1016/j.cell.2020.02.052. - DOI - PMC - PubMed
-
- Zhai X., Sun J., Yan Z., Zhang J., Zhao J., Zhao Z., Gao Q., He W.-T., Veit M., Su S. Comparison of Severe Acute Respiratory Syndrome Coronavirus 2 Spike Protein Binding to ACE2 Receptors from Human, Pets, Farm Animals, and Putative Intermediate Hosts. J. Virol. 2020;94:e00831-20. doi: 10.1128/JVI.00831-20. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

