A synergy between the catalytic and structural Zn(II) ions and the enzyme and substrate dynamics underlies the structure-function relationships of matrix metalloproteinase collagenolysis
- PMID: 34228191
- PMCID: PMC9018075
- DOI: 10.1007/s00775-021-01876-6
A synergy between the catalytic and structural Zn(II) ions and the enzyme and substrate dynamics underlies the structure-function relationships of matrix metalloproteinase collagenolysis
Abstract
Matrix metalloproteinases (MMPs) are Zn(II) dependent endopeptidases involved in the degradation of collagen. Unbalanced collagen breakdown results in numerous pathological conditions, including cardiovascular and neurodegenerative diseases and tumor growth and invasion. Matrix metalloproteinase-1 (MMP-1) is a member of the MMPs family. The enzyme contains catalytic and structural Zn(II) ions. Despite many studies on the enzyme, there is little known about the synergy between the two Zn(II) metal ions and the enzyme and substrate dynamics in MMP-1 structure-function relationships. We performed a computational study of the MMP-1•triple-helical peptide (THP) enzyme•substrate complex to provide this missing insight. Our results revealed Zn(II) ions' importance in modulating the long-range correlated motions in the MMP-1•THP complex. Overall, our results reveal the importance of the catalytic Zn(II) and the role of the structural Zn(II) ion in preserving the integrity of the enzyme active site and the overall enzyme-substrate complex synergy with the dynamics of the enzyme and the substrate. Notably, both Zn(II) sites participate in diverse networks of long-range correlated motions that involve the CAT and HPX domains and the THP substrate, thus exercising a complex role in the stability and functionality of the MMP-1•THP complex. Both the Zn(II) ions have a distinct impact on the structural stability and dynamics of the MMP-1•THP complex. The study shifts the paradigm from the "local role" of the Zn(II) ions with knowledge about their essential role in the long-range dynamics and stability of the overall enzyme•substrate (ES) complex.
Keywords: Catalytic and structural zinc centers; Computational modeling; Matrix metalloproteinase-1; Molecular dynamics; Zinc dependent enzymes.
© 2021. Society for Biological Inorganic Chemistry (SBIC).
Figures
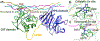
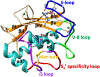

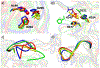
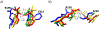
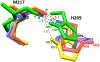


References
-
- Piez KA, Eigner EA, and Lewis MS (1963) The Chromatographic Separation and Amino Acid Composition of the Subunits of Several Collagens. Biochemistry 2:58–66
-
- Gallop PM, Blumenfield OO, Seifter S (1972) Structure and Metabolism of Connective tissue proteins. Ann Rev Biochem 41:617–672 - PubMed
-
- Miller EJ (1971) Isolation and Characterization of the Cyanogen Bromide Peptides from the α1(II) Chain of Chick Cartilage Collagen. Biochemistry 10:3030–3034 - PubMed
-
- Trelstad RL, Kang AH, Toole BP, Gross J (1972) Collagen Heterogeneity: High resolution separation of native [α1(I)]2α2 and [α1(II)]3 and their component α chains. J Biol Chem 247:6469–6473 - PubMed
-
- Chung E, Miller EJ (1974) Collagen Polymorphism: Characterization of Molecules with the Chain Composition [α1(III)]3 in Human Tissues. Science 183:1200–1201 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

