FANCD2-Associated Nuclease 1 Partially Compensates for the Lack of Exonuclease 1 in Mismatch Repair
- PMID: 34228493
- PMCID: PMC8384067
- DOI: 10.1128/MCB.00303-21
FANCD2-Associated Nuclease 1 Partially Compensates for the Lack of Exonuclease 1 in Mismatch Repair
Abstract
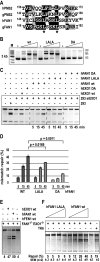
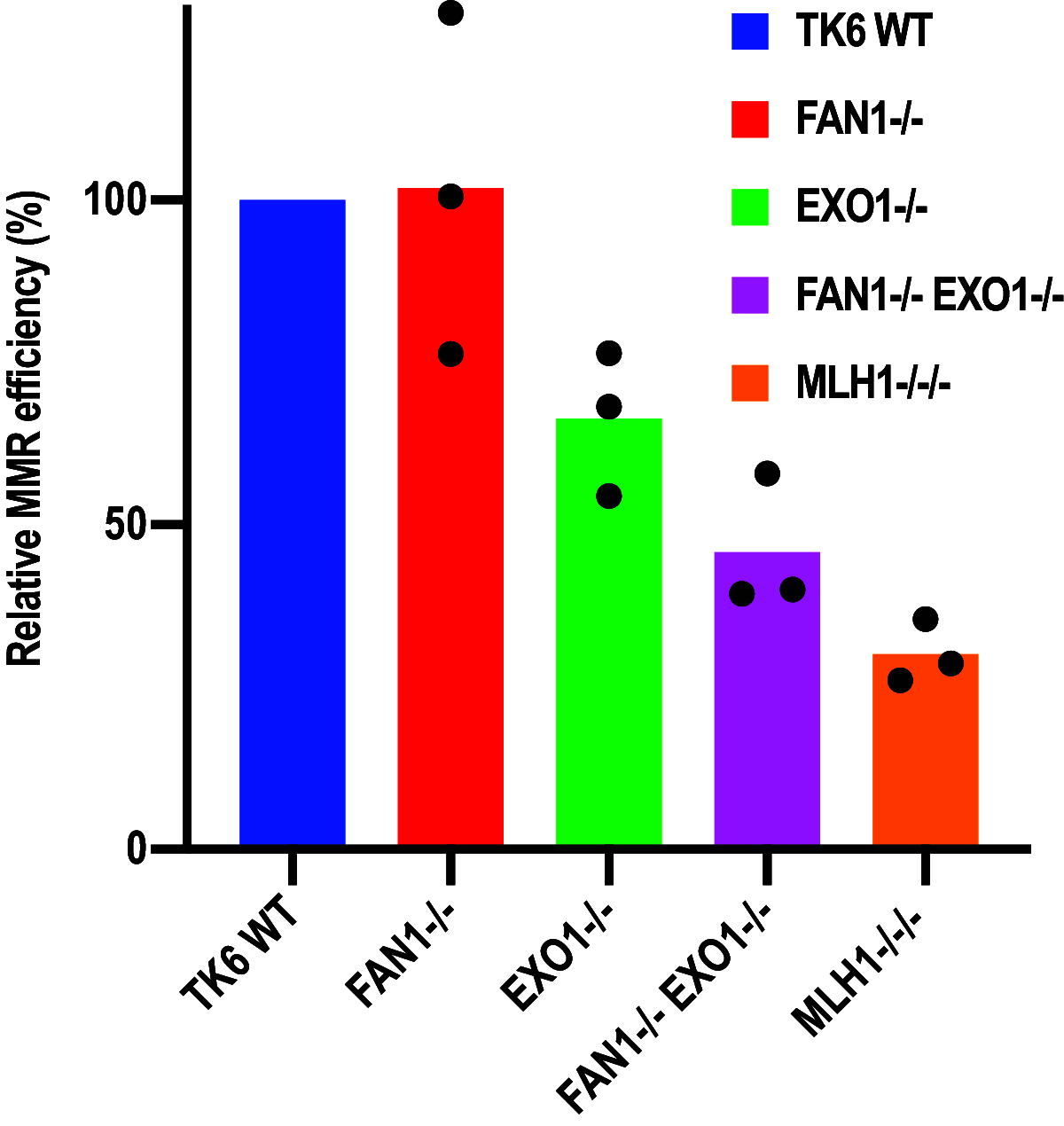
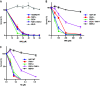
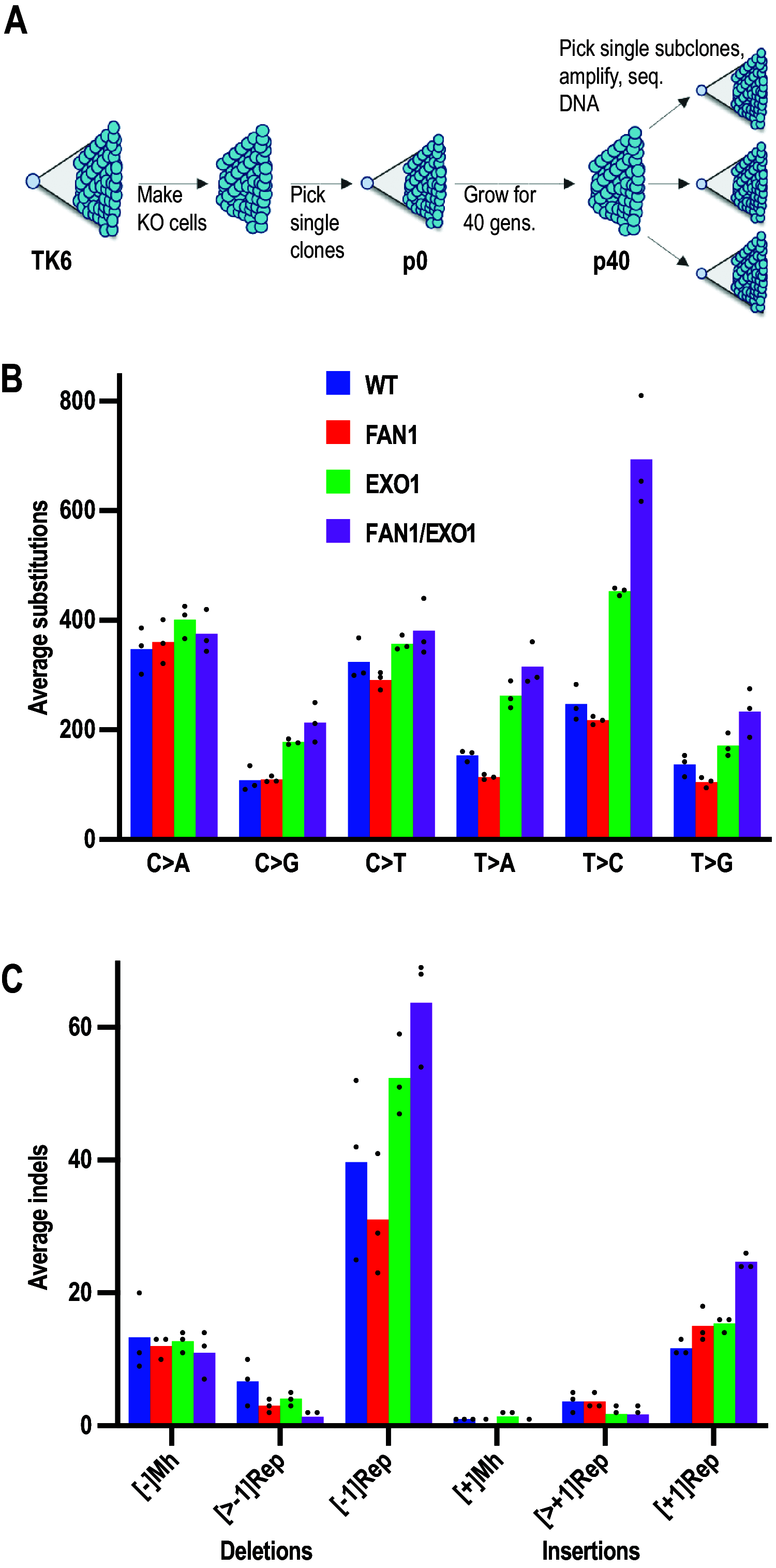
Germline mutations in the mismatch repair (MMR) genes MSH2, MSH6, MLH1, and PMS2 are linked to cancer of the colon and other organs, characterized by microsatellite instability and a large increase in mutation frequency. Unexpectedly, mutations in EXO1, encoding the only exonuclease genetically implicated in MMR, are not linked to familial cancer and cause a substantially weaker mutator phenotype. This difference could be explained if eukaryotic cells possessed additional exonucleases redundant with EXO1. Analysis of the MLH1 interactome identified FANCD2-associated nuclease 1 (FAN1), a novel enzyme with biochemical properties resembling EXO1. We now show that FAN1 efficiently substitutes for EXO1 in MMR assays and that this functional complementation is modulated by its interaction with MLH1. FAN1 also contributes to MMR in vivo; cells lacking both EXO1 and FAN1 have an MMR defect and display resistance to N-methyl-N-nitrosourea (MNU) and 6-thioguanine (TG). Moreover, FAN1 loss amplifies the mutational profile of EXO1-deficient cells, suggesting that the two nucleases act redundantly in the same antimutagenic pathway. However, the increased drug resistance and mutator phenotype of FAN1/EXO1-deficient cells are less prominent than those seen in cells lacking MSH6 or MLH1. Eukaryotic cells thus apparently possess additional mechanisms that compensate for the loss of EXO1.
Keywords: DNA repair; EXO1; FAN1; MLH1; MSH6; exonuclease; mismatch repair; mutational signature.
Figures

Similar articles
-
FAN1's protection against CGG repeat expansion requires its nuclease activity and is FANCD2-independent.Nucleic Acids Res. 2021 Nov 18;49(20):11643-11652. doi: 10.1093/nar/gkab899. Nucleic Acids Res. 2021. PMID: 34718701 Free PMC article.
-
MLH1-mediated recruitment of FAN1 to chromatin for the induction of apoptosis triggered by O6 -methylguanine.Genes Cells. 2020 Mar;25(3):175-186. doi: 10.1111/gtc.12748. Epub 2020 Feb 6. Genes Cells. 2020. PMID: 31955481
-
Exo1 independent DNA mismatch repair involves multiple compensatory nucleases.DNA Repair (Amst). 2014 Sep;21:55-64. doi: 10.1016/j.dnarep.2014.06.005. Epub 2014 Jul 15. DNA Repair (Amst). 2014. PMID: 25037770 Free PMC article.
-
Exonuclease 1-dependent and independent mismatch repair.DNA Repair (Amst). 2015 Aug;32:24-32. doi: 10.1016/j.dnarep.2015.04.010. Epub 2015 Apr 30. DNA Repair (Amst). 2015. PMID: 25956862 Free PMC article. Review.
-
Exonuclease 1 and its versatile roles in DNA repair.Crit Rev Biochem Mol Biol. 2016 Nov/Dec;51(6):440-451. doi: 10.1080/10409238.2016.1215407. Epub 2016 Aug 5. Crit Rev Biochem Mol Biol. 2016. PMID: 27494243 Review.
Cited by
-
FAN1's protection against CGG repeat expansion requires its nuclease activity and is FANCD2-independent.Nucleic Acids Res. 2021 Nov 18;49(20):11643-11652. doi: 10.1093/nar/gkab899. Nucleic Acids Res. 2021. PMID: 34718701 Free PMC article.
-
A CRISPR-Cas9 screen identifies EXO1 as a formaldehyde resistance gene.Nat Commun. 2023 Jan 24;14(1):381. doi: 10.1038/s41467-023-35802-y. Nat Commun. 2023. PMID: 36693839 Free PMC article.
-
The nuclease activity of DNA2 promotes exonuclease 1-independent mismatch repair.J Biol Chem. 2022 Apr;298(4):101831. doi: 10.1016/j.jbc.2022.101831. Epub 2022 Mar 15. J Biol Chem. 2022. PMID: 35300981 Free PMC article.
-
FAN1 removes triplet repeat extrusions via a PCNA- and RFC-dependent mechanism.Proc Natl Acad Sci U S A. 2023 Aug 15;120(33):e2302103120. doi: 10.1073/pnas.2302103120. Epub 2023 Aug 7. Proc Natl Acad Sci U S A. 2023. PMID: 37549289 Free PMC article.
-
Double strand breaks drive toxicity in Huntington's disease mice with or without somatic expansion.bioRxiv [Preprint]. 2025 May 28:2025.05.27.654663. doi: 10.1101/2025.05.27.654663. bioRxiv. 2025. PMID: 40501854 Free PMC article. Preprint.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
