Multicolor Whole-Genome Mapping in Nanochannels for Genetic Analysis
- PMID: 34232611
- PMCID: PMC9705121
- DOI: 10.1021/acs.analchem.1c01373
Multicolor Whole-Genome Mapping in Nanochannels for Genetic Analysis
Abstract
Analysis of structural variations (SVs) is important to understand mutations underlying genetic disorders and pathogenic conditions. However, characterizing SVs using short-read, high-throughput sequencing technology is difficult. Although long-read sequencing technologies are being increasingly employed in characterizing SVs, their low throughput and high costs discourage widespread adoption. Sequence motif-based optical mapping in nanochannels is useful in whole-genome mapping and SV detection, but it is not possible to precisely locate the breakpoints or estimate the copy numbers. We present here a universal multicolor mapping strategy in nanochannels combining conventional sequence-motif labeling system with Cas9-mediated target-specific labeling of any 20-base sequences (20mers) to create custom labels and detect new features. The sequence motifs are labeled with green fluorophores and the 20mers are labeled with red fluorophores. Using this strategy, it is possible to not only detect the SVs but also utilize custom labels to interrogate the features not accessible to motif-labeling, locate breakpoints, and precisely estimate copy numbers of genomic repeats. We validated our approach by quantifying the D4Z4 copy numbers, a known biomarker for facioscapulohumeral muscular dystrophy (FSHD) and estimating the telomere length, a clinical biomarker for assessing disease risk factors in aging-related diseases and malignant cancers. We also demonstrate the application of our methodology in discovering transposable long non-interspersed Elements 1 (LINE-1) insertions across the whole genome.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
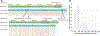


References
-
- Macintyre G; Ylstra B; Brenton JD Trends Genet. 2016, 32, 530–542. - PubMed
-
- Sleegers K; Brouwers N; Gijselinck I; Theuns J; Goossens D; Wauters J; Del-Favero J; Cruts M; Duijn C. M. v.; Broeckhoven CV Brain 2006, 129, 2977–2983. - PubMed
-
- Rovelet-Lecrux A; Hannequin D; Raux G; Meur NL; Laquerrière A; Vital A; Dumanchin C; Feuillette S; Brice A; Vercelletto M; Dubas F; Frebourg T; Campion D Nat. Genet. 2006, 38, 24–26. - PubMed
-
- Talkowski ME; Rosenfeld JA; Blumenthal I; Pillalamarri V; Chiang C; Heilbut A; Ernst C; Hanscom C; Rossin E; Lindgren AM; Pereira S; Ruderfer D; Kirby A; Ripke S; Harris DJ; Lee J-H; Ha K; Kim H-G; Solomon BD; Gropman AL; Lucente D; Sims K; Ohsumi TK; Borowsky ML; Loranger S; Quade B; Lage K; Miles J; Wu B-L; Shen Y; Neale B; Shaffer LG; Daly MJ; Morton CC; Gusella JF Cell 2012, 149, 525–537. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

