Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer's disease
- PMID: 34239132
- PMCID: PMC8766217
- DOI: 10.1038/s41588-021-00894-z
Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer's disease
Abstract
The gene-regulatory landscape of the brain is highly dynamic in health and disease, coordinating a menagerie of biological processes across distinct cell types. Here, we present a multi-omic single-nucleus study of 191,890 nuclei in late-stage Alzheimer's disease (AD), accessible through our web portal, profiling chromatin accessibility and gene expression in the same biological samples and uncovering vast cellular heterogeneity. We identified cell-type-specific, disease-associated candidate cis-regulatory elements and their candidate target genes, including an oligodendrocyte-associated regulatory module containing links to APOE and CLU. We describe cis-regulatory relationships in specific cell types at a subset of AD risk loci defined by genome-wide association studies, demonstrating the utility of this multi-omic single-nucleus approach. Trajectory analysis of glial populations identified disease-relevant transcription factors, such as SREBF1, and their regulatory targets. Finally, we introduce single-nucleus consensus weighted gene coexpression analysis, a coexpression network analysis strategy robust to sparse single-cell data, and perform a systems-level analysis of the AD transcriptome.
© 2021. The Author(s), under exclusive licence to Springer Nature America, Inc.
Figures



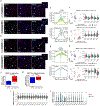














Similar articles
-
Sex dependent glial-specific changes in the chromatin accessibility landscape in late-onset Alzheimer's disease brains.Mol Neurodegener. 2021 Aug 24;16(1):58. doi: 10.1186/s13024-021-00481-0. Mol Neurodegener. 2021. PMID: 34429139 Free PMC article.
-
Developmental dynamics of the single nucleus regulatory landscape of pig hippocampus.Sci China Life Sci. 2023 Nov;66(11):2614-2628. doi: 10.1007/s11427-022-2345-2. Epub 2023 Jul 6. Sci China Life Sci. 2023. PMID: 37428306
-
Single-Cell Transcriptomic Profiling Reveals Regional Differences in the Prefrontal and Entorhinal Cortex of Alzheimer's Disease Brain.Int J Mol Sci. 2025 May 19;26(10):4841. doi: 10.3390/ijms26104841. Int J Mol Sci. 2025. PMID: 40429980 Free PMC article.
-
Profiling Chromatin Accessibility at Single-cell Resolution.Genomics Proteomics Bioinformatics. 2021 Apr;19(2):172-190. doi: 10.1016/j.gpb.2020.06.010. Epub 2021 Feb 11. Genomics Proteomics Bioinformatics. 2021. PMID: 33581341 Free PMC article. Review.
-
Characterizing cis-regulatory elements using single-cell epigenomics.Nat Rev Genet. 2023 Jan;24(1):21-43. doi: 10.1038/s41576-022-00509-1. Epub 2022 Jul 15. Nat Rev Genet. 2023. PMID: 35840754 Free PMC article. Review.
Cited by
-
Nuclei Detection and Segmentation of Histopathological Images Using a Feature Pyramidal Network Variant of a Mask R-CNN.Bioengineering (Basel). 2024 Oct 1;11(10):994. doi: 10.3390/bioengineering11100994. Bioengineering (Basel). 2024. PMID: 39451370 Free PMC article.
-
Identification and validation of key biomarkers associated with macrophages in nonalcoholic fatty liver disease based on hdWGCNA and machine learning.Aging (Albany NY). 2023 Dec 21;15(24):15451-15472. doi: 10.18632/aging.205374. Epub 2023 Dec 21. Aging (Albany NY). 2023. PMID: 38147020 Free PMC article.
-
Deciphering the impact of aging on splenic endothelial cell heterogeneity and immunosenescence through single-cell RNA sequencing analysis.Immun Ageing. 2024 Jul 18;21(1):48. doi: 10.1186/s12979-024-00452-1. Immun Ageing. 2024. PMID: 39026350 Free PMC article.
-
The chromatin accessibility and transcriptomic landscape of the aging mice cochlea and the identification of potential functional super-enhancers in age-related hearing loss.Clin Epigenetics. 2024 Jul 4;16(1):86. doi: 10.1186/s13148-024-01702-1. Clin Epigenetics. 2024. PMID: 38965562 Free PMC article.
-
Human cerebral organoids - a new tool for clinical neurology research.Nat Rev Neurol. 2022 Nov;18(11):661-680. doi: 10.1038/s41582-022-00723-9. Epub 2022 Oct 17. Nat Rev Neurol. 2022. PMID: 36253568 Free PMC article. Review.
References
Methods only references
-
- Stuart T, Srivastava A, Lareau C & Satija R Multimodal single-cell chromatin analysis with Signac. bioRxiv 2020.11.09.373613 (2020).
-
- Gu Z, Eils R & Schlesner M Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

