Direct RT-qPCR assay for SARS-CoV-2 variants of concern (Alpha, B.1.1.7 and Beta, B.1.351) detection and quantification in wastewater
- PMID: 34245731
- PMCID: PMC8262398
- DOI: 10.1016/j.envres.2021.111653
Direct RT-qPCR assay for SARS-CoV-2 variants of concern (Alpha, B.1.1.7 and Beta, B.1.351) detection and quantification in wastewater
Abstract
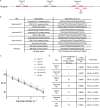
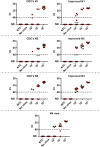
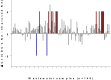
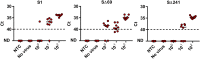
Less than a year following the Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) outbreak, variants of concern have emerged in the form of variant Alpha (B.1.1.7, the British variant) and Beta (B.1.351, the South Africa variant). Due to their high infectivity and morbidity, it has become clear that it is crucial to quickly and effectively detect these and other variants. Here, we report improved primers-probe sets for reverse transcriptase quantitative polymerase chain reaction (RT-qPCR) for SARS-CoV-2 detection including a rapid, cost-effective, and direct RT-qPCR method for detection of the two variants of concern (Alpha, B.1.1.7 and Beta, B.1.351). All the developed primers-probe sets were fully characterized, demonstrating sensitive and specific detection. These primer-probe sets were also successfully employed on wastewater samples aimed at detecting and even quantifying new variants in a geographical area, even prior to the reports by the medical testing. The novel primers-probe sets presented here will enable proper responses for pandemic containment, particularly considering the emergence of variants of concern.
Keywords: Molecular probes; Reverse transcriptase polymerase chain reaction; SARS-CoV-2; Variants of concern; Wastewater-based epidemiological monitoring.
Copyright © 2021 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

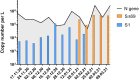
References
-
- Ahmed W., Angel N., Edson J., Bibby K., Bivins A., O'Brien J.W., Choi P.M., Kitajima M., Simpson S.L., Li J., Tscharke B., Verhagen R., Smith W.J.M., Zaugg J., Dierens L., Hugenholtz P., Thomas K.V., Mueller J.F. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020;728:138764. doi: 10.1016/j.scitotenv.2020.138764. - DOI - PMC - PubMed
-
- Ahmed W., Bivins A., Bertsch P.M., Bibby K., Choi P.M., Farkas K., Gyawali P., Hamilton K.A., Haramoto E., Kitajima M., Simpson S.L., Tandukar S., Thomas K.V., Mueller J.F. Surveillance of SARS-CoV-2 RNA in wastewater: methods optimization and quality control are crucial for generating reliable public health information. Curr. Opin. Environ. Sci. Heal. 2020;17:82–93. doi: 10.1016/j.coesh.2020.09.003. - DOI - PMC - PubMed
-
- Andrés C., Garcia-Cehic D., Gregori J., Piñana M., Rodriguez-Frias F., Guerrero-Murillo M., Esperalba J., Rando A., Goterris L., Codina M.G., Quer S., Martín M.C., Campins M., Ferrer R., Almirante B., Esteban J.I., Pumarola T., Antón A., Quer J. Naturally occurring SARS-CoV-2 gene deletions close to the spike S1/S2 cleavage site in the viral quasispecies of COVID19 patients. Emerg. Microb. Infect. 2020;9:1900–1911. doi: 10.1080/22221751.2020.1806735. - DOI - PMC - PubMed
-
- Bar-Or I., Yaniv K., Shagan M., Ozer E., Erster O., Mendelson E., Mannasse B., Shirazi R., Kramarsky-Winter E., Nir O., Abu-Ali H., Ronen Z., Rinott E., Lewis Y.E., Friedler E., Bitkover E., Paitan Y., Berchenko Y., Kushmaro A. Regressing SARS-CoV-2 sewage measurements onto COVID-19 burden in the population: a proof-of-concept for quantitative environmental surveillance. 2020. medRxiv. 1–11. - DOI - PMC - PubMed
-
- Brown J.C., Goldhill D.H., Zhou J., Peacock T.P., Frise R., Goonawardane N., Baillon L., Kugathasan R., Pinto A.L., McKay P.F., Hassard J., Moshe M., Singanayagam A., Burgoyne T., Investigators the A. Consortium P.V., Barclay W.S. Increased transmission of SARS-CoV-2 lineage B . 1 . 1 . 7 ( VOC 2020212/01 ) is not accounted for by a replicative advantage in primary airway cells or antibody escape. 2021. bioRxiv. 1–13. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

