Structure, Application, and Biochemistry of Microbial Keratinases
- PMID: 34248885
- PMCID: PMC8260994
- DOI: 10.3389/fmicb.2021.674345
Structure, Application, and Biochemistry of Microbial Keratinases
Abstract
Keratinases belong to a class of proteases that are able to degrade keratins into amino acids. Microbial keratinases play important roles in turning keratin-containing wastes into value-added products by participating in the degradation of keratin. Keratin is found in human and animal hard tissues, and its complicated structures make it resistant to degradation by common proteases. Although breaking disulfide bonds are involved in keratin degradation, keratinase is responsible for the cleavage of peptides, making it attractive in pharmaceutical and feather industries. Keratinase can serve as an important tool to convert keratin-rich wastes such as feathers from poultry industry into diverse products applicable to many fields. Despite of some progress made in isolating keratinase-producing microorganisms, structural studies of keratinases, and biochemical characterization of these enzymes, effort is still required to expand the biotechnological application of keratinase in diverse fields by identifying more keratinases, understanding the mechanism of action and constructing more active enzymes through molecular biology and protein engineering. Herein, this review covers structures, applications, biochemistry of microbial keratinases, and strategies to improve its efficiency in keratin degradation.
Keywords: keratin; keratinase; microorganisms; protease; structure; waste treatment.
Copyright © 2021 Li.
Conflict of interest statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



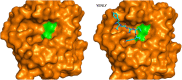

Similar articles
-
Progress in Microbial Degradation of Feather Waste.Front Microbiol. 2019 Dec 5;10:2717. doi: 10.3389/fmicb.2019.02717. eCollection 2019. Front Microbiol. 2019. PMID: 31866957 Free PMC article. Review.
-
Comprehensive insights into microbial keratinases and their implication in various biotechnological and industrial sectors: A review.Int J Biol Macromol. 2020 Jul 1;154:567-583. doi: 10.1016/j.ijbiomac.2020.03.116. Epub 2020 Mar 16. Int J Biol Macromol. 2020. PMID: 32194110 Review.
-
Molecular strategies to enhance the keratinase gene expression and its potential implications in poultry feed industry.Poult Sci. 2024 May;103(5):103606. doi: 10.1016/j.psj.2024.103606. Epub 2024 Mar 5. Poult Sci. 2024. PMID: 38479096 Free PMC article.
-
Microbial Keratinases: Enzymes with Promising Biotechnological Applications.Food Technol Biotechnol. 2018 Sep;56(3):312-328. doi: 10.17113/ftb.56.03.18.5658. Food Technol Biotechnol. 2018. PMID: 30510475 Free PMC article. Review.
-
Microbial enzymes catalyzing keratin degradation: Classification, structure, function.Biotechnol Adv. 2020 Nov 15;44:107607. doi: 10.1016/j.biotechadv.2020.107607. Epub 2020 Aug 5. Biotechnol Adv. 2020. PMID: 32768519 Free PMC article. Review.
Cited by
-
Chicken Feather Waste Valorization Into Nutritive Protein Hydrolysate: Role of Novel Thermostable Keratinase From Bacillus pacificus RSA27.Front Microbiol. 2022 Apr 25;13:882902. doi: 10.3389/fmicb.2022.882902. eCollection 2022. Front Microbiol. 2022. PMID: 35547122 Free PMC article.
-
Genome-wide analysis of Keratinibaculum paraultunense strain KD-1 T and its key genes and metabolic pathways involved in the anaerobic degradation of feather keratin.Arch Microbiol. 2022 Sep 20;204(10):634. doi: 10.1007/s00203-022-03226-9. Arch Microbiol. 2022. PMID: 36127480
-
Alkaline keratinase from Bacillus sp. DRS4 efficiently biodegrades chicken feathers to synthesize improved keratin/bacterial nanocellulose-based bioplastics.Heliyon. 2024 Jun 9;10(12):e32768. doi: 10.1016/j.heliyon.2024.e32768. eCollection 2024 Jun 30. Heliyon. 2024. PMID: 38975182 Free PMC article.
-
Comprehensive insights into the mechanism of keratin degradation and exploitation of keratinase to enhance the bioaccessibility of soybean protein.Biotechnol Biofuels Bioprod. 2023 Nov 17;16(1):177. doi: 10.1186/s13068-023-02426-9. Biotechnol Biofuels Bioprod. 2023. PMID: 37978558 Free PMC article.
-
Keratinous bioresources: their generation, microbial degradation, and value enhancement for biotechnological applications.World J Microbiol Biotechnol. 2025 Mar 29;41(4):118. doi: 10.1007/s11274-025-04336-4. World J Microbiol Biotechnol. 2025. PMID: 40155538 Review.
References
-
- Anbu P., Hilda A., Sur H.-W., Hur B.-K., Jayanthi S. (2008). Extracellular keratinase from Trichophyton sp. HA-2 isolated from feather dumping soil. Int. Biodeterior. Biodegradation 62 287–292. 10.1016/j.ibiod.2007.07.017 - DOI
-
- Awad G. E. A., Esawy M. A., Salam W. A., Salama B. M., Abdelkader A. F., El-diwany A. (2011). Keratinase production by Bacillus pumilus GHD in solid-state fermentation using sugar cane bagasse: optimisation of culture conditions using a Box-Behnken experimental design. Ann. Microbiol. 61 663–672. 10.1007/s13213-010-0187-0 - DOI
-
- Bach E., Sant’Anna V., Daroit D. J., Corrêa A. P. F., Segalin J., Brandelli A. (2012). Production, one-step purification, and characterization of a keratinolytic protease from Serratia marcescens P3. Process Biochem. 47 2455–2462. 10.1016/j.procbio.2012.10.007 - DOI
Publication types
LinkOut - more resources
Full Text Sources

