Virus hunters: Discovering the evolutionary origins of SARS-CoV-2
- PMID: 34265239
- PMCID: PMC8279507
- DOI: 10.1016/j.chom.2021.06.012
Virus hunters: Discovering the evolutionary origins of SARS-CoV-2
Abstract
The likely animal source of SARS-CoV-2 remains speculative. A recent study published in Cell by Zhou et al. reported the detection of novel alpha- and betacoronaviruses, including SARS-CoV-2-related viruses in bats.
Copyright © 2021 Elsevier Inc. All rights reserved.
Figures
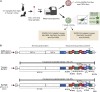
Comment on
-
Identification of novel bat coronaviruses sheds light on the evolutionary origins of SARS-CoV-2 and related viruses.Cell. 2021 Aug 19;184(17):4380-4391.e14. doi: 10.1016/j.cell.2021.06.008. Epub 2021 Jun 9. Cell. 2021. PMID: 34147139 Free PMC article.
References
-
- Drosten C., Günther S., Preiser W., van der Werf S., Brodt H.R., Becker S., Rabenau H., Panning M., Kolesnikova L., Fouchier R.A., et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003;348:1967–1976. - PubMed
-
- Guan Y., Zheng B.J., He Y.Q., Liu X.L., Zhuang Z.X., Cheung C.L., Luo S.W., Li P.H., Zhang L.J., Guan Y.J., et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science. 2003;302:276–278. - PubMed
-
- Li W., Shi Z., Yu M., Ren W., Smith C., Epstein J.H., Wang H., Crameri G., Hu Z., Zhang H., et al. Bats are natural reservoirs of SARS-like coronaviruses. Science. 2005;310:676–679. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

