Cortex cis-regulatory switches establish scale colour identity and pattern diversity in Heliconius
- PMID: 34280087
- PMCID: PMC8289415
- DOI: 10.7554/eLife.68549
Cortex cis-regulatory switches establish scale colour identity and pattern diversity in Heliconius
Abstract
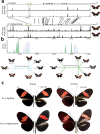
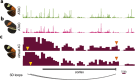
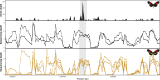
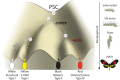
In Heliconius butterflies, wing colour pattern diversity and scale types are controlled by a few genes of large effect that regulate colour pattern switches between morphs and species across a large mimetic radiation. One of these genes, cortex, has been repeatedly associated with colour pattern evolution in butterflies. Here we carried out CRISPR knockouts in multiple Heliconius species and show that cortex is a major determinant of scale cell identity. Chromatin accessibility profiling and introgression scans identified cis-regulatory regions associated with discrete phenotypic switches. CRISPR perturbation of these regions in black hindwing genotypes recreated a yellow bar, revealing their spatially limited activity. In the H. melpomene/timareta lineage, the candidate CRE from yellow-barred phenotype morphs is interrupted by a transposable element, suggesting that cis-regulatory structural variation underlies these mimetic adaptations. Our work shows that cortex functionally controls scale colour fate and that its cis-regulatory regions control a phenotypic switch in a modular and pattern-specific fashion.
Keywords: ATAC-Seq; cis-regulation; crisppr; developmental biology; evolution; evolutionary biology; heliconius; wing patterning.
Plain language summary
Heliconius butterflies have bright patterns on their wings that tell potential predators that they are toxic. As a result, predators learn to avoid eating them. Over time, unrelated species of butterflies have evolved similar patterns to avoid predation through a process known as Müllerian mimicry. Worldwide, there are over 180,000 species of butterflies and moths, most of which have different wing patterns. How do genes create this pattern diversity? And do butterflies use similar genes to create similar wing patterns? One of the genes involved in creating wing patterns is called cortex. This gene has a large region of DNA around it that does not code for proteins, but instead, controls whether cortex is on or off in different parts of the wing. Changes in this non-coding region can act like switches, turning regions of the wing into different colours and creating complex patterns, but it is unclear how these switches have evolved. Butterfly wings get their colour from tiny structures called scales, which each have their own unique set of pigments. In Heliconius butterflies, there are three types of scales: yellow/white scales, black scales, and red/orange/brown scales. Livraghi et al. used a DNA editing technique called CRISPR to find out whether the cortex gene affects scale type. First, Livraghi et al. confirmed that deleting cortex turned black and red scales yellow. Next, they used the same technique to manipulate the non-coding DNA around the cortex gene to see the effect on the wing pattern. This manipulation turned a black-winged butterfly into a butterfly with a yellow wing band, a pattern that occurs naturally in Heliconius butterflies. The next step was to find the mutation responsible for the appearance of yellow wing bands in nature. It turns out that a bit of extra genetic code, derived from so-called ‘jumping genes’, had inserted itself into the non-coding DNA around the cortex gene, ‘flipping’ the switch and leading to the appearance of the yellow scales. Genetic information contains the instructions to generate shape and form in most organisms. These instructions evolve over millions of years, creating everything from bacteria to blue whales. Butterfly wings are visual evidence of evolution, but the way their genes create new patterns isn't specific to butterflies. Understanding wing patterns can help researchers to learn how genetic switches control diversity across other species too.
© 2021, Livraghi et al.
Conflict of interest statement
LL, JH, SV, GM, Ev, LL, AR, IW, JL, CC, LH, CW, JW, JF, ZG, HA, CS, MP, RP, AM, WM, CJ No competing interests declared
Figures

































Similar articles
-
Aristaless Controls Butterfly Wing Color Variation Used in Mimicry and Mate Choice.Curr Biol. 2018 Nov 5;28(21):3469-3474.e4. doi: 10.1016/j.cub.2018.08.051. Epub 2018 Oct 25. Curr Biol. 2018. PMID: 30415702 Free PMC article.
-
Interplay between Developmental Flexibility and Determinism in the Evolution of Mimetic Heliconius Wing Patterns.Curr Biol. 2019 Dec 2;29(23):3996-4009.e4. doi: 10.1016/j.cub.2019.10.010. Epub 2019 Nov 14. Curr Biol. 2019. PMID: 31735676
-
Perfect mimicry between Heliconius butterflies is constrained by genetics and development.Proc Biol Sci. 2020 Jul 29;287(1931):20201267. doi: 10.1098/rspb.2020.1267. Epub 2020 Jul 22. Proc Biol Sci. 2020. PMID: 32693728 Free PMC article.
-
Waiting in the wings: what can we learn about gene co-option from the diversification of butterfly wing patterns?Philos Trans R Soc Lond B Biol Sci. 2017 Feb 5;372(1713):20150485. doi: 10.1098/rstb.2015.0485. Philos Trans R Soc Lond B Biol Sci. 2017. PMID: 27994126 Free PMC article. Review.
-
Genes controlling mimetic colour pattern variation in butterflies.Curr Opin Insect Sci. 2016 Oct;17:24-31. doi: 10.1016/j.cois.2016.05.013. Epub 2016 May 27. Curr Opin Insect Sci. 2016. PMID: 27720070 Review.
Cited by
-
Evolutionary Changes in the Chromatin Landscape Contribute to Reorganization of a Developmental Gene Network During Rapid Life History Evolution in Sea Urchins.Mol Biol Evol. 2022 Sep 1;39(9):msac172. doi: 10.1093/molbev/msac172. Mol Biol Evol. 2022. PMID: 35946348 Free PMC article.
-
A butterfly pan-genome reveals that a large amount of structural variation underlies the evolution of chromatin accessibility.Genome Res. 2022 Oct;32(10):1862-1875. doi: 10.1101/gr.276839.122. Epub 2022 Sep 15. Genome Res. 2022. PMID: 36109150 Free PMC article.
-
aristaless1 has a dual role in appendage formation and wing color specification during butterfly development.BMC Biol. 2023 May 4;21(1):100. doi: 10.1186/s12915-023-01601-6. BMC Biol. 2023. PMID: 37143075 Free PMC article.
-
A large deletion at the cortex locus eliminates butterfly wing patterning.G3 (Bethesda). 2022 Apr 4;12(4):jkac021. doi: 10.1093/g3journal/jkac021. G3 (Bethesda). 2022. PMID: 35099556 Free PMC article.
-
Surprise hybrid origins of a butterfly species.Nature. 2024 Apr;628(8009):723-724. doi: 10.1038/d41586-024-00858-3. Nature. 2024. PMID: 38632416 No abstract available.
References
-
- Aymone AC, Valente VL, de Araújo AM. Ultrastructure and morphogenesis of the wing scales in Heliconius erato Phyllis (Lepidoptera: nymphalidae): what silvery/brownish surfaces can tell Us about the development of color patterning? Arthropod Structure & Development. 2013;42:349–359. doi: 10.1016/j.asd.2013.06.001. - DOI - PubMed
-
- Brien MN, Enciso-Romero J, Parnell AJ, Salazar PA, Morochz C, Chalá D, Bainbridge HE, Zinn T, Curran EV, Nadeau NJ. Phenotypic variation in Heliconius erato crosses shows that iridescent structural colour is sex-linked and controlled by multiple genes. Interface Focus. 2019;9:20180047. doi: 10.1098/rsfs.2018.0047. - DOI - PMC - PubMed
-
- Brown KS. The biology of Heliconius and related genera. Annual Review of Entomology. 1981;26:427–457. doi: 10.1146/annurev.en.26.010181.002235. - DOI

