Engineered Type Six Secretion Systems Deliver Active Exogenous Effectors and Cre Recombinase
- PMID: 34281388
- PMCID: PMC8406194
- DOI: 10.1128/mBio.01115-21
Engineered Type Six Secretion Systems Deliver Active Exogenous Effectors and Cre Recombinase
Abstract
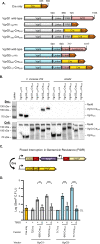
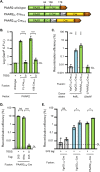
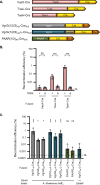
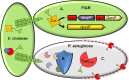
Genetic editing has revolutionized biotechnology, but delivery of endonuclease genes as DNA can lead to aberrant integration or overexpression, leading to off-target effects. Here, we develop a mechanism to deliver Cre recombinase as a protein by engineering the bacterial type six secretion system (T6SS). Using multiple T6SS fusion proteins, Aeromonas dhakensis or attenuated Vibrio cholerae donor strains, and a gain-of-function cassette for detecting Cre recombination, we demonstrate successful delivery of active Cre directly into recipient cells. The most efficient transfer was achieved using a truncated version of PAAR2 from V. cholerae, resulting in a relatively small (118-amino-acid) delivery tag. We further demonstrate the versatility of this system by delivering an exogenous effector, TseC, enabling V. cholerae to kill Pseudomonas aeruginosa. This implies that P. aeruginosa is naturally resistant to all native effectors of V. cholerae and that the TseC chaperone protein is not required for its activity. Moreover, it demonstrates that the engineered system can improve T6SS efficacy against specific pathogens, proposing future application in microbiome manipulation or as a next-generation antimicrobial. Inexpensive and easy to produce, this protein delivery system has many potential applications, ranging from studying T6SS effectors to genetic editing. IMPORTANCE Delivery of protein-based drugs, antigens, and gene-editing agents has broad applications. The type VI protein secretion system (T6SS) can target both bacteria and eukaryotic cells and deliver proteins of diverse size and function. Here, we harness the T6SS to successfully deliver Cre recombinase to genetically edit bacteria without requiring the introduction of exogenous DNA into the recipient cells. This demonstrates a promising advantage over current genetic editing tools that require transformation or conjugation of DNA. The engineered secretion tag can also deliver a heterologous antimicrobial toxin that kills an otherwise unsusceptible pathogen, Pseudomonas aeruginosa. These results demonstrate the potential of T6SS-mediated delivery in areas including genome editing, killing drug-resistant pathogens, and studying toxin functions.
Keywords: Aeromonas dhakensis; Cre recombinase; DNA recombination; Pseudomonas aeruginosa; T6SS; Vibrio cholerae; antimicrobial; bactericidal activity; biotechnology; effector; genetic editing; interspecies interactions; protein engineering; protein secretion; type six secretion system.
Figures

Similar articles
-
Pesticin-Like Effector VgrG3cp Targeting Peptidoglycan Delivered by the Type VI Secretion System Contributes to Vibrio cholerae Interbacterial Competition.Microbiol Spectr. 2023 Feb 14;11(1):e0426722. doi: 10.1128/spectrum.04267-22. Epub 2023 Jan 10. Microbiol Spectr. 2023. PMID: 36625646 Free PMC article.
-
Characterization of Lysozyme-Like Effector TseP Reveals the Dependence of Type VI Secretion System (T6SS) Secretion on Effectors in Aeromonas dhakensis Strain SSU.Appl Environ Microbiol. 2021 May 26;87(12):e0043521. doi: 10.1128/AEM.00435-21. Epub 2021 May 26. Appl Environ Microbiol. 2021. PMID: 33837015 Free PMC article.
-
A Dueling-Competent Signal-Sensing Module Guides Precise Delivery of Cargo Proteins into Target Cells by Engineered Pseudomonas aeruginosa.ACS Synth Biol. 2023 Feb 17;12(2):360-368. doi: 10.1021/acssynbio.2c00441. Epub 2023 Jan 20. ACS Synth Biol. 2023. PMID: 36662232
-
The Vibrio cholerae type VI secretion system: toxins, regulators and consequences.Environ Microbiol. 2020 Oct;22(10):4112-4122. doi: 10.1111/1462-2920.14976. Epub 2020 Mar 13. Environ Microbiol. 2020. PMID: 32133757 Review.
-
Type VI secretion system effector proteins: Effective weapons for bacterial competitiveness.Cell Microbiol. 2020 Sep;22(9):e13241. doi: 10.1111/cmi.13241. Epub 2020 Jul 19. Cell Microbiol. 2020. PMID: 32592518 Review.
Cited by
-
Type VI Secretion Systems: Environmental and Intra-host Competition of Vibrio cholerae.Adv Exp Med Biol. 2023;1404:41-63. doi: 10.1007/978-3-031-22997-8_3. Adv Exp Med Biol. 2023. PMID: 36792870
-
Emerging methylation-based approaches in microbiome engineering.Biotechnol Biofuels Bioprod. 2024 Jul 10;17(1):96. doi: 10.1186/s13068-024-02529-x. Biotechnol Biofuels Bioprod. 2024. PMID: 38987811 Free PMC article. Review.
-
Global biogeography and projection of antimicrobial toxin genes.Microbiome. 2025 Feb 4;13(1):40. doi: 10.1186/s40168-025-02038-5. Microbiome. 2025. PMID: 39905479 Free PMC article.
-
Not just passengers: effectors contribute to the assembly of the type VI secretion system as structural building blocks.J Bacteriol. 2025 Mar 20;207(3):e0045524. doi: 10.1128/jb.00455-24. Epub 2025 Feb 4. J Bacteriol. 2025. PMID: 39902958 Free PMC article. Review.
-
Recruitment of heterologous substrates by bacterial secretion systems for transkingdom translocation.Front Cell Infect Microbiol. 2023 Mar 6;13:1146000. doi: 10.3389/fcimb.2023.1146000. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 36949816 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

