Archaea Are Rare and Uncommon Members of the Mammalian Skin Microbiome
- PMID: 34282942
- PMCID: PMC8407491
- DOI: 10.1128/mSystems.00642-21
Archaea Are Rare and Uncommon Members of the Mammalian Skin Microbiome
Abstract
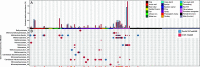
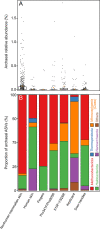
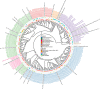
Although previous research demonstrates that skin-associated archaea are rarely detected within human skin microbiome data, exist at relatively low abundance, and are primarily affiliated with the Methanobacteriota and Halobacteriota phyla, other studies suggest that archaea are consistently detected and relatively abundant on human skin, with skin "archaeomes" dominated by putative ammonia oxidizers of the Nitrososphaeria class (Thermoproteota phylum, formerly Thaumarchaeota). Here, we evaluated new and existing 16S rRNA gene sequence data sourced from mammalian skin and skin-associated surfaces and generated with two commonly used universal prokaryotic primer sets to assess archaeal prevalence, relative abundance, and taxonomic distribution. Archaeal 16S rRNA gene sequences were detected in only 17.5% of 1,688 samples by high-throughput sequence data, with most of the archaeon-positive samples associated with nonhuman mammalian skin. Only 5.9% of human-associated skin sample data sets contained sequences affiliated with archaeal 16S rRNA genes. When detected, the relative abundance of sequences affiliated with archaeal amplicon sequence variants (ASVs) was less than 1% for most mammalian skin samples and did not exceed 2% for any samples. Although several computer keyboard microbial profiles were dominated by Nitrososphaeria sequences, all other skin microbiome data sets tested were primarily composed of sequences affiliated with Methanobacteriota and Halobacteriota phyla. Our findings revise downward recent estimates of human skin archaeal distributions and relative abundances, especially those affiliated with the Nitrososphaeria, reflecting a limited and infrequent archaeal presence within the mammalian skin microbiome. IMPORTANCE The current state of research on mammalian skin-associated archaea is limited, with the few papers focusing on potential skin archaeal communities often in disagreement with each other. As such, there is no consensus on the prevalence or taxonomic composition of archaea on mammalian skin. Mammalian skin health is in part influenced by its complex microbiota and consortium of bacteria and potential archaea. Without a clear foundational analysis and characterization of the mammalian skin archaeome, it will be difficult for future research to explore the potential impact of skin-associated archaea on skin health and function. The current work provides a much-needed analysis of the mammalian skin archaeome and contributes to building a foundation from which further discussion and exploration of the skin archaeome might continue.
Keywords: 16S RNA; archaea; built environment; gene sequencing; mammalian skin; microbiome; rare biosphere.
Figures

References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

