Extended-spectrum β-lactamases: an update on their characteristics, epidemiology and detection
- PMID: 34286272
- PMCID: PMC8284625
- DOI: 10.1093/jacamr/dlab092
Extended-spectrum β-lactamases: an update on their characteristics, epidemiology and detection
Abstract
Extended-spectrum β-lactamase (ESBL)-producing Gram-negative pathogens are a major cause of resistance to expanded-spectrum β-lactam antibiotics. Since their discovery in the early 1980s, they have spread worldwide and an are now endemic in Enterobacterales isolated from both hospital-associated and community-acquired infections. As a result, they are a global public health concern. In the past, TEM- and SHV-type ESBLs were the predominant families of ESBLs. Today CTX-M-type enzymes are the most commonly found ESBL type with the CTX-M-15 variant dominating worldwide, followed in prevalence by CTX-M-14, and CTX-M-27 is emerging in certain parts of the world. The genes encoding ESBLs are often found on plasmids and harboured within transposons or insertion sequences, which has enabled their spread. In addition, the population of ESBL-producing Escherichia coli is dominated globally by a highly virulent and successful clone belonging to ST131. Today, there are many diagnostic tools available to the clinical microbiology laboratory and include both phenotypic and genotypic tests to detect β-lactamases. Unfortunately, when ESBLs are not identified in a timely manner, appropriate antimicrobial therapy is frequently delayed, resulting in poor clinical outcomes. Several analyses of clinical trials have shown mixed results with regards to whether a carbapenem must be used to treat serious infections caused by ESBLs or whether some of the older β-lactam-β-lactamase combinations such as piperacillin/tazobactam are appropriate. Some of the newer combinations such as ceftazidime/avibactam have demonstrated efficacy in patients. ESBL-producing Gram-negative pathogens will continue to be major contributor to antimicrobial resistance worldwide. It is essential that we remain vigilant about identifying them both in patient isolates and through surveillance studies.
© The Author(s) 2021. Published by Oxford University Press on behalf of the British Society for Antimicrobial Chemotherapy.
Figures
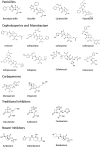
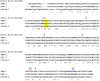


References
-
- Bush K, Fisher JF.. Epidemiological expansion, structural studies, and clinical challenges of new β-lactamases from gram-negative bacteria. Annu Rev Microbiol 2011; 65: 455–78. - PubMed
-
- Pitout JD, Nordmann P, Laupland KB. et al. Emergence of Enterobacteriaceae producing extended-spectrum b-lactamases (ESBLs) in the community. J Antimicrob Chemother 2005; 56: 52–9. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
