A Multistep Workflow to Evaluate Newly Generated iPSCs and Their Ability to Generate Different Cell Types
- PMID: 34287353
- PMCID: PMC8293472
- DOI: 10.3390/mps4030050
A Multistep Workflow to Evaluate Newly Generated iPSCs and Their Ability to Generate Different Cell Types
Abstract
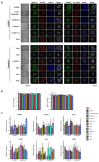
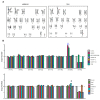
Induced pluripotent stem cells (iPSCs) derived from human somatic cells have created new opportunities to generate disease-relevant cells. Thus, as the use of patient-derived stem cells has become more widespread, having a workflow to monitor each line is critical. This ensures iPSCs pass a suite of quality-control measures, promoting reproducibility across experiments and between labs. With this in mind, we established a multistep workflow to assess our newly generated iPSCs. Our workflow tests four benchmarks: cell growth, genomic stability, pluripotency, and the ability to form the three germline layers. We also outline a simple test for assessing cell growth and highlight the need to compare different growth media. Genomic integrity in the human iPSCs is analyzed by G-band karyotyping and a qPCR-based test for the detection of common karyotypic abnormalities. Finally, we confirm that the iPSC lines can differentiate into a given cell type, using a trilineage assay, and later confirm that each iPSC can be differentiated into one cell type of interest, with a focus on the generation of cortical neurons. Taken together, we present a multistep quality-control workflow to evaluate newly generated iPSCs and detail the findings on these lines as they are tested within the workflow.
Keywords: cortical neurons; genomic integrity; human-induced pluripotent stem cells; neural progenitor cells; quality control; trilineage differentiation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

