Correlated Motions in Structural Biology
- PMID: 34291898
- PMCID: PMC9059239
- DOI: 10.1021/acs.biochem.1c00420
Correlated Motions in Structural Biology
Abstract
Correlated motions in proteins arising from the collective movements of residues have long been proposed to be fundamentally important to key properties of proteins, from allostery and catalysis to evolvability. Recent breakthroughs in structural biology have made it possible to capture proteins undergoing complex conformational changes, yet intrinsic correlated motions within a conformation remain one of the least understood facets of protein structure. For many decades, the analysis of total X-ray scattering held the promise of animating crystal structures with correlated motions. With recent advances in both X-ray detectors and data interpretation methods, this long-held promise can now be met. In this Perspective, we will introduce how correlated motions are captured in total scattering and provide guidelines for the collection, interpretation, and validation of data. As structural biology continues to push the boundaries, we see an opportunity to gain atomistic insight into correlated motions using total scattering as a bridge between theory and experiment.
Figures
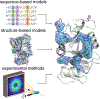



References
-
- Kühlbrandt W (2014) The resolution revolution. Science 343, 1443–1444. - PubMed
-
- Reich ES (2013) Ultimate upgrade for US synchrotron. Nature 501, 148–149. - PubMed
-
- Hand E (2009) X-ray free-electron lasers fire up. Nature 461, 708–709. - PubMed
-
- Chapman HN, Fromme P, Barty A, White TA, Kirian RA, Aquila A, Hunter MS, Schulz J, Deponte DP, Weierstall U, Doak RB, Maia FRNC, Martin AV, Schlichting I, Lomb L, Coppola N, Shoeman RL, Epp SW, Hartmann R, Rolles D, Rudenko A, Foucar L, Kimmel N, Weidenspointner G, Holl P, Liang M, Barthelmess M, Caleman C, Boutet S, Bogan MJ, Krzywinski J, Bostedt C, Bajt S, Gumprecht L, Rudek B, Erk B, Schmidt C, Hömke A, Reich C, Pietschner D, Ströder L, Hauser G, Gorke H, Ullrich J, Herrmann S, Schaller G, Schopper F, Soltau H, Kühnel KU, Messerschmidt M, Bozek JD, Hau-Riege SP, Frank M, Hampton CY, Sierra RG, Starodub D, Williams GJ, Hajdu J, Timneanu N, Seibert MM, Andreasson J, Rocker A, Jönsson O, Svenda M, Stern S, Nass K, Andritschke R, Schröter CD, Krasniqi F, Bott M, Schmidt KE, Wang X, Grotjohann I, Holton JM, Barends TRM, Neutze R, Marchesini S, Fromme R, Schorb S, Rupp D, Adolph M, Gorkhover T, Andersson I, Hirsemann H, Potdevin G, Graafsma H, Nilsson B, and Spence JCH (2011) Femtosecond X-ray protein nanocrystallography. Nature 470, 73–78. - PMC - PubMed
-
- Stellato F, Oberthür D, Liang M, Bean R, Gati C, Yefanov O, Barty A, Burkhardt A, Fischer P, Galli L, Kirian RA, Meyer J, Panneerselvam S, Yoon CH, Chervinskii F, Speller E, White TA, Betzel C, Meents A, and Chapman HN (2014) Room-temperature macromolecular serial crystallography using synchrotron radiation. IUCrJ 1, 204–212. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

