Adjacent Cell Marker Lateral Spillover Compensation and Reinforcement for Multiplexed Images
- PMID: 34295327
- PMCID: PMC8289709
- DOI: 10.3389/fimmu.2021.652631
Adjacent Cell Marker Lateral Spillover Compensation and Reinforcement for Multiplexed Images
Abstract
Multiplex imaging technologies are now routinely capable of measuring more than 40 antibody-labeled parameters in single cells. However, lateral spillage of signals in densely packed tissues presents an obstacle to the assignment of high-dimensional spatial features to individual cells for accurate cell-type annotation. We devised a method to correct for lateral spillage of cell surface markers between adjacent cells termed REinforcement Dynamic Spillover EliminAtion (REDSEA). The use of REDSEA decreased contaminating signals from neighboring cells. It improved the recovery of marker signals across both isotopic (i.e., Multiplexed Ion Beam Imaging) and immunofluorescent (i.e., Cyclic Immunofluorescence) multiplexed images resulting in a marked improvement in cell-type classification.
Keywords: cell annotation; image correction; multiplexed tissue imaging; signal spillover; single-cell biology; spatial proteomics.
Copyright © 2021 Bai, Zhu, Rovira-Clave, Chen, Markovic, Chan, Su, McIlwain, Estes, Keren, Nolan and Jiang.
Conflict of interest statement
GN is a co-founder and has a personal financial interest in the company IonPath, which manufactures the instrument used in this manuscript. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

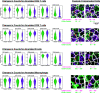
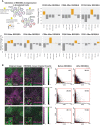

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

