New binding specificities evolve via point mutation in an invertebrate allorecognition gene
- PMID: 34296075
- PMCID: PMC8282982
- DOI: 10.1016/j.isci.2021.102811
New binding specificities evolve via point mutation in an invertebrate allorecognition gene
Abstract
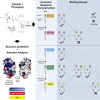
Many organisms use genetic self-recognition systems to distinguish themselves from conspecifics. In the cnidarian, Hydractinia symbiolongicarpus, self-recognition is partially controlled by allorecognition 2 (Alr2). Alr2 encodes a highly polymorphic transmembrane protein that discriminates self from nonself by binding in trans to other Alr2 proteins with identical or similar sequences. Here, we focused on the N-terminal domain of Alr2, which can determine its binding specificity. We pair ancestral sequence reconstruction and experimental assays to show that amino acid substitutions can create sequences with novel binding specificities either directly (via one mutation) or via sequential mutations and intermediates with relaxed specificities. We also show that one side of the domain has experienced positive selection and likely forms the binding interface. Our results provide direct evidence that point mutations can generate Alr2 proteins with novel binding specificities. This provides a plausible mechanism for the generation and maintenance of functional variation in nature.
Keywords: Evolutionary Biology; Molecular Biology; Molecular Genetics.
© 2021 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
A family of unusual immunoglobulin superfamily genes in an invertebrate histocompatibility complex.Proc Natl Acad Sci U S A. 2022 Oct 4;119(40):e2207374119. doi: 10.1073/pnas.2207374119. Epub 2022 Sep 26. Proc Natl Acad Sci U S A. 2022. PMID: 36161920 Free PMC article.
-
Allorecognition proteins in an invertebrate exhibit homophilic interactions.Curr Biol. 2015 Nov 2;25(21):2845-2850. doi: 10.1016/j.cub.2015.09.030. Epub 2015 Oct 8. Curr Biol. 2015. PMID: 26455308 Free PMC article.
-
The Hydractinia allorecognition system.Immunogenetics. 2022 Feb;74(1):27-34. doi: 10.1007/s00251-021-01233-6. Epub 2021 Nov 13. Immunogenetics. 2022. PMID: 34773127 Review.
-
EVOLUTIONARY GENETICS OF ALLORECOGNITION IN THE COLONIAL HYDROID HYDRACTINIA SYMBIOLONGICARPUS.Evolution. 1996 Dec;50(6):2221-2240. doi: 10.1111/j.1558-5646.1996.tb03612.x. Evolution. 1996. PMID: 28565652
-
Model systems of invertebrate allorecognition.Curr Biol. 2011 Jan 25;21(2):R82-92. doi: 10.1016/j.cub.2010.11.061. Curr Biol. 2011. PMID: 21256442 Review.
Cited by
-
A family of unusual immunoglobulin superfamily genes in an invertebrate histocompatibility complex.Proc Natl Acad Sci U S A. 2022 Oct 4;119(40):e2207374119. doi: 10.1073/pnas.2207374119. Epub 2022 Sep 26. Proc Natl Acad Sci U S A. 2022. PMID: 36161920 Free PMC article.
References
-
- Abràmoff M.D., Magalhães P.J., Ram S.J. Image processing with ImageJ. Biophotonics Int. 2004;7:36–42.
-
- Buss L.W. Princeton University Press; 1987. The Evolution of Individuality.
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

