Hedgehog/GLI Signaling Pathway: Transduction, Regulation, and Implications for Disease
- PMID: 34298625
- PMCID: PMC8304605
- DOI: 10.3390/cancers13143410
Hedgehog/GLI Signaling Pathway: Transduction, Regulation, and Implications for Disease
Abstract
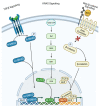
The Hh/GLI signaling pathway was originally discovered in Drosophila as a major regulator of segment patterning in development. This pathway consists of a series of ligands (Shh, Ihh, and Dhh), transmembrane receptors (Ptch1 and Ptch2), transcription factors (GLI1-3), and signaling regulators (SMO, HHIP, SUFU, PKA, CK1, GSK3β, etc.) that work in concert to repress (Ptch1, Ptch2, SUFU, PKA, CK1, GSK3β) or activate (Shh, Ihh, Dhh, SMO, GLI1-3) the signaling cascade. Not long after the initial discovery, dysregulation of the Hh/GLI signaling pathway was implicated in human disease. Activation of this signaling pathway is observed in many types of cancer, including basal cell carcinoma, medulloblastoma, colorectal, prostate, pancreatic, and many more. Most often, the activation of the Hh/GLI pathway in cancer occurs through a ligand-independent mechanism. However, in benign disease, this activation is mostly ligand-dependent. The upstream signaling component of the receptor complex, SMO, is bypassed, and the GLI family of transcription factors can be activated regardless of ligand binding. Additional mechanisms of pathway activation exist whereby the entirety of the downstream signaling pathway is bypassed, and PTCH1 promotes cell cycle progression and prevents caspase-mediated apoptosis. Throughout this review, we summarize each component of the signaling cascade, non-canonical modes of pathway activation, and the implications in human disease, including cancer.
Keywords: GLI; Hh pathway inhibitors; SUFU; cancer; canonical and non-canonical activation; hedgehog.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Lindsley D.L., Grell E.H. Genetic Variations of Drosophila Melanogaster. 627th ed. Carnegie Institution of Washington Publication; Washington, DC, USA: 1983.
-
- Ekker S., Jackson D., Von Kessler D., Sun B., Young K., Beachy P., Ekker S., Jackson D., Von Kessler D., Sun B., et al. The degree of variation in DNA sequence recognition among four Drosophila homeotic proteins. EMBO J. 1994;13:3551–3560. doi: 10.1002/j.1460-2075.1994.tb06662.x. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

