Structural basis for ligand binding modes of CTP synthase
- PMID: 34301892
- PMCID: PMC8325340
- DOI: 10.1073/pnas.2026621118
Structural basis for ligand binding modes of CTP synthase
Abstract
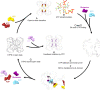
Cytidine triphosphate synthase (CTPS), which comprises an ammonia ligase domain and a glutamine amidotransferase domain, catalyzes the final step of de novo CTP biosynthesis. The activity of CTPS is regulated by the binding of four nucleotides and glutamine. While glutamine serves as an ammonia donor for the ATP-dependent conversion of UTP to CTP, the fourth nucleotide GTP acts as an allosteric activator. Models have been proposed to explain the mechanisms of action at the active site of the ammonia ligase domain and the conformational changes derived by GTP binding. However, actual GTP/ATP/UTP binding modes and relevant conformational changes have not been revealed fully. Here, we report the discovery of binding modes of four nucleotides and a glutamine analog 6-diazo-5-oxo-L-norleucine in Drosophila CTPS by cryo-electron microscopy with near-atomic resolution. Interactions between GTP and surrounding residues indicate that GTP acts to coordinate reactions at both domains by directly blocking ammonia leakage and stabilizing the ammonia tunnel. Additionally, we observe the ATP-dependent UTP phosphorylation intermediate and determine interacting residues at the ammonia ligase. A noncanonical CTP binding at the ATP binding site suggests another layer of feedback inhibition. Our findings not only delineate the structure of CTPS in the presence of all substrates but also complete our understanding of the underlying mechanisms of the allosteric regulation and CTP synthesis.
Keywords: CTP synthase; allosteric regulation; cryo–electron microscopy; cytoophidium.
Copyright © 2021 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures





Similar articles
-
GTP-Dependent Regulation of CTP Synthase: Evolving Insights into Allosteric Activation and NH3 Translocation.Biomolecules. 2022 Apr 29;12(5):647. doi: 10.3390/biom12050647. Biomolecules. 2022. PMID: 35625575 Free PMC article. Review.
-
"Pinching" the ammonia tunnel of CTP synthase unveils coordinated catalytic and allosteric-dependent control of ammonia passage.Biochim Biophys Acta Gen Subj. 2018 Dec;1862(12):2714-2727. doi: 10.1016/j.bbagen.2018.08.008. Epub 2018 Aug 10. Biochim Biophys Acta Gen Subj. 2018. PMID: 30251661
-
Competition between ammonia derived from internal glutamine hydrolysis and hydroxylamine present in the solution for incorporation into UTP as catalysed by Lactococcus lactis CTP synthase.Arch Biochem Biophys. 2004 Apr 1;424(1):105-11. doi: 10.1016/j.abb.2004.01.018. Arch Biochem Biophys. 2004. PMID: 15019842
-
Inhibition of E. coli CTP synthase by the "positive" allosteric effector GTP.Biochim Biophys Acta. 2004 Jun 1;1699(1-2):213-20. doi: 10.1016/j.bbapap.2004.03.002. Biochim Biophys Acta. 2004. PMID: 15158730
-
CTPS cytoophidia in Drosophila: distribution, regulation, and physiological roles.Exp Cell Res. 2025 Apr 15;447(2):114536. doi: 10.1016/j.yexcr.2025.114536. Epub 2025 Mar 22. Exp Cell Res. 2025. PMID: 40122502 Review.
Cited by
-
Combined Inactivation of CTPS1 and ATR Is Synthetically Lethal to MYC-Overexpressing Cancer Cells.Cancer Res. 2022 Mar 15;82(6):1013-1024. doi: 10.1158/0008-5472.CAN-21-1707. Cancer Res. 2022. PMID: 35022212 Free PMC article.
-
The role of filamentation in activation and DNA sequence specificity of the sequence-specific endonuclease SgrAI.Biochem Soc Trans. 2022 Dec 16;50(6):1703-1714. doi: 10.1042/BST20220547. Biochem Soc Trans. 2022. PMID: 36398769 Free PMC article.
-
Structural basis of dynamic P5CS filaments.Elife. 2022 Mar 14;11:e76107. doi: 10.7554/eLife.76107. Elife. 2022. PMID: 35286254 Free PMC article.
-
Fat body-specific reduction of CTPS alleviates HFD-induced obesity.Elife. 2023 Sep 11;12:e85293. doi: 10.7554/eLife.85293. Elife. 2023. PMID: 37695169 Free PMC article.
-
Structural basis for isoform-specific inhibition of human CTPS1.Proc Natl Acad Sci U S A. 2021 Oct 5;118(40):e2107968118. doi: 10.1073/pnas.2107968118. Proc Natl Acad Sci U S A. 2021. PMID: 34583994 Free PMC article.
References
-
- Kizaki H., Williams J. C., Morris H. P., Weber G., Increased cytidine 5′-triphosphate synthetase activity in rat and human tumors. Cancer Res. 40, 3921–3927 (1980). - PubMed
-
- Williams J. C., Kizaki H., Weber G., Morris H. P., Increased CTP synthetase activity in cancer cells. Nature 271, 71–73 (1978). - PubMed
-
- Ellims P. H., Gan T. E., Medley G., Cytidine triphosphate synthetase activity in lymphoproliferative disorders. Cancer Res. 43, 1432–1435 (1983). - PubMed
-
- De Clercq E., Antiviral agents: Characteristic activity spectrum depending on the molecular target with which they interact. Adv. Virus Res. 42, 1–55 (1993). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

