Modelling of gene loss propensity in the pangenomes of three Brassica species suggests different mechanisms between polyploids and diploids
- PMID: 34310022
- PMCID: PMC8633514
- DOI: 10.1111/pbi.13674
Modelling of gene loss propensity in the pangenomes of three Brassica species suggests different mechanisms between polyploids and diploids
Abstract
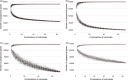
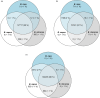
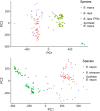
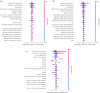
Plant genomes demonstrate significant presence/absence variation (PAV) within a species; however, the factors that lead to this variation have not been studied systematically in Brassica across diploids and polyploids. Here, we developed pangenomes of polyploid Brassica napus and its two diploid progenitor genomes B. rapa and B. oleracea to infer how PAV may differ between diploids and polyploids. Modelling of gene loss suggests that loss propensity is primarily associated with transposable elements in the diploids while in B. napus, gene loss propensity is associated with homoeologous recombination. We use these results to gain insights into the different causes of gene loss, both in diploids and following polyploidization, and pave the way for the application of machine learning methods to understanding the underlying biological and physical causes of gene presence/absence.
Keywords: Brassica; XGBoost; gene loss propensity; machine learning; pangenome; transposable elements.
© 2021 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interests.
Figures
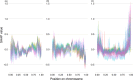
References
-
- Alexa, A. and Rahnenführer, J. (2009) Gene set enrichment analysis with topGO. Bioconductor Improve, 27, 1–26.
-
- Alix, K. , Joets, J. , Ryder, C.D. , Moore, J. , Barker, G.C. , Bailey, J.P. , King, G.J. et al. (2008) The CACTA transposon Bot1 played a major role in Brassica genome divergence and gene proliferation. Plant J. 56, 1030–1044. - PubMed
-
- Allainguillaume, J. , Alexander, M. , Bullock, J. , Saunders, M. , Allender, C.J. , King, G. , Ford, C.S. et al. (2006) Fitness of hybrids between rapeseed (Brassica napus) and wild Brassica rapa in natural habitats. Mol. Ecol. 15, 1175–1184. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

