Janus Bioparticles: Asymmetric Nucleosomes and Their Preparation Using Chemical Biology Approaches
- PMID: 34319695
- PMCID: PMC8411803
- DOI: 10.1021/acs.accounts.1c00313
Janus Bioparticles: Asymmetric Nucleosomes and Their Preparation Using Chemical Biology Approaches
Abstract
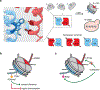
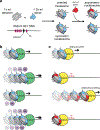
The fundamental repeating unit of chromatin, the nucleosome, is composed of DNA wrapped around two copies each of four canonical histone proteins. Nucleosomes possess 2-fold pseudo-symmetry that is subject to disruption in cellular contexts. For example, the post-translational modification (PTM) of histones plays an essential role in epigenetic regulation, and the introduction of a PTM on only one of the two "sister" histone copies in a given nucleosome eliminates the inherent symmetry of the complex. Similarly, the removal or swapping of histones for variants or the introduction of a histone mutant may render the two faces of the nucleosome asymmetric, creating, if you will, a type of "Janus" bioparticle. Over the past decade, many groups have detailed the discovery of asymmetric species in chromatin isolated from numerous cell types. However, in vitro biochemical and biophysical investigation of asymmetric nucleosomes has proven synthetically challenging. Whereas symmetric nucleosomes are readily formed via a stochastic combination of their histone and DNA components, asymmetric nucleosome assembly demands the selective incorporation of a single modified/mutant histone copy alongside its wild-type counterpart.Herein we describe the chemical biology tools that we and others have developed in recent years for investigating nucleosome asymmetry. Such approaches, each with its own benefits and shortcomings, fall into five broad categories. First, we discuss affinity tag-based purification methods. These enable the assembly of theoretically any asymmetric nucleosome of interest but are frequently labor-intensive and suffer from low yields. Second, we detail transient cross-linking strategies that are amenable to the preparation of histone H3- or H4-modified/mutant asymmetric species. These yield asymmetric nucleosomes in a traceless fashion, albeit through the use of more complicated synthesis techniques. Third, we describe a synthetic biology technique based on the generation of bump-hole mutant H3 histones that selectively heterodimerize. Although currently developed only for H3 and a related isoform, this method uniquely allows for the interrogation of nucleosome asymmetry in yeast. Fourth, we outline a method for generating H2A- or H2B-modified/mutant asymmetric nucleosomes that relies on the differential DNA-histone contact strength inherent in the Widom 601 DNA sequence. This technique involves the initial formation of hexasomes which are then complemented with distinct H2A/H2B dimers. Finally, we review an approach that utilizes split intein technology to isolate asymmetric H2A- or H2B-modified/mutant nucleosomes. This method shares steps in common with the former but exploits tagged, intein-fused dimers for the facile purification of asymmetric products.Throughout the Account, we highlight various biological questions that drove the development of these methods and ultimately were answered by them. Though each technique has its own shortcomings, collectively these chemical biology tools provide a means to biochemically interrogate a plethora of asymmetric nucleosome species. We conclude with a discussion of remaining challenges, particularly that of endogenous asymmetric nucleosome detection.
Conflict of interest statement
The authors declare no competing financial interest.
Figures




Similar articles
-
Synthesis of Oriented Hexasomes and Asymmetric Nucleosomes Using a Template Editing Process.J Am Chem Soc. 2022 Feb 9;144(5):2284-2291. doi: 10.1021/jacs.1c12420. Epub 2022 Jan 26. J Am Chem Soc. 2022. PMID: 35081309 Free PMC article.
-
Structural analysis of the hexasome, lacking one histone H2A/H2B dimer from the conventional nucleosome.Biochemistry. 2012 Apr 17;51(15):3302-9. doi: 10.1021/bi300129b. Epub 2012 Apr 2. Biochemistry. 2012. PMID: 22448809
-
Nucleosome adaptability conferred by sequence and structural variations in histone H2A-H2B dimers.Curr Opin Struct Biol. 2015 Jun;32:48-57. doi: 10.1016/j.sbi.2015.02.004. Epub 2015 Feb 27. Curr Opin Struct Biol. 2015. PMID: 25731851 Free PMC article. Review.
-
H2A-H2B Histone Dimer Plasticity and Its Functional Implications.Cells. 2022 Sep 12;11(18):2837. doi: 10.3390/cells11182837. Cells. 2022. PMID: 36139412 Free PMC article.
-
Nucleosomal asymmetry: a novel mechanism to regulate nucleosome function.Biochem Soc Trans. 2024 Jun 26;52(3):1219-1232. doi: 10.1042/BST20230877. Biochem Soc Trans. 2024. PMID: 38778762 Free PMC article. Review.
Cited by
-
Structure and dynamics of the chromatin remodeler ALC1 bound to a PARylated nucleosome.Elife. 2021 Sep 6;10:e71420. doi: 10.7554/eLife.71420. Elife. 2021. PMID: 34486521 Free PMC article.
-
Synthesis of Oriented Hexasomes and Asymmetric Nucleosomes Using a Template Editing Process.J Am Chem Soc. 2022 Feb 9;144(5):2284-2291. doi: 10.1021/jacs.1c12420. Epub 2022 Jan 26. J Am Chem Soc. 2022. PMID: 35081309 Free PMC article.
-
NLRP3 Inflammasome-Mediated Osteoarthritis: The Role of Epigenetics.Biology (Basel). 2025 Jan 14;14(1):71. doi: 10.3390/biology14010071. Biology (Basel). 2025. PMID: 39857301 Free PMC article. Review.
-
A circular engineered sortase for interrogating histone H3 in chromatin.bioRxiv [Preprint]. 2024 Sep 10:2024.09.10.612318. doi: 10.1101/2024.09.10.612318. bioRxiv. 2024. Update in: J Am Chem Soc. 2024 Dec 11;146(49):33914-33927. doi: 10.1021/jacs.4c12585. PMID: 39372790 Free PMC article. Updated. Preprint.
-
The spread of chemical biology into chromatin.J Biol Chem. 2024 Nov;300(11):107776. doi: 10.1016/j.jbc.2024.107776. Epub 2024 Sep 12. J Biol Chem. 2024. PMID: 39276931 Free PMC article. Review.
References
-
- Dao HT; Dul BE; Dann GP; Liszczak GP; Muir TW, A basic motif anchoring ISWI to nucleosome acidic patch regulates nucleosome spacing. Nature Chemical Biology 2020, 16, 134–142. - PMC - PubMed
-
Application of the oriented hexasome-based assembly method to study the effects of asymmetric acidic patch mutations on ISWI-mediated chromatin remodeling.
-
- Bagert JD; Mitchener MM; Patriotis AL; Dul BE; Wojcik F; Nacev BA; Feng L; Allis CD; Muir TW, Oncohistone mutations enhance chromatin remodeling and alter cell fates. Nature Chemical Biology 2021, 17, 403–411. - PMC - PubMed
-
Introduction of the tagged split-intein method for formation of asymmetric nucleosomes.
-
- Luger K; Mader AW; Richmond RK; Sargent DF; Richmond TJ, Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 1997, 389, 251–260. - PubMed
-
- Allis CD; Jenuwein T, The molecular hallmarks of epigenetic control. Nature Reviews Genetics 2016, 17, 487–500. - PubMed
-
- Strahl BD; Allis CD, The language of covalent histone modifications. Nature 2000, 403, 41–45. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

