Altered neuronal physiology, development, and function associated with a common chromosome 15 duplication involving CHRNA7
- PMID: 34320968
- PMCID: PMC8317352
- DOI: 10.1186/s12915-021-01080-7
Altered neuronal physiology, development, and function associated with a common chromosome 15 duplication involving CHRNA7
Abstract
Background: Copy number variants (CNVs) linked to genes involved in nervous system development or function are often associated with neuropsychiatric disease. While CNVs involving deletions generally cause severe and highly penetrant patient phenotypes, CNVs leading to duplications tend instead to exhibit widely variable and less penetrant phenotypic expressivity among affected individuals. CNVs located on chromosome 15q13.3 affecting the alpha-7 nicotinic acetylcholine receptor subunit (CHRNA7) gene contribute to multiple neuropsychiatric disorders with highly variable penetrance. However, the basis of such differential penetrance remains uncharacterized. Here, we generated induced pluripotent stem cell (iPSC) models from first-degree relatives with a 15q13.3 duplication and analyzed their cellular phenotypes to uncover a basis for the dissimilar phenotypic expressivity.
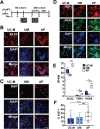
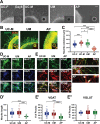
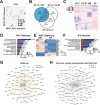
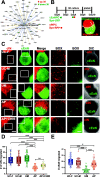
Results: The first-degree relatives studied included a boy with autism and emotional dysregulation (the affected proband-AP) and his clinically unaffected mother (UM), with comparison to unrelated control models lacking this duplication. Potential contributors to neuropsychiatric impairment were modeled in iPSC-derived cortical excitatory and inhibitory neurons. The AP-derived model uniquely exhibited disruptions of cellular physiology and neurodevelopment not observed in either the UM or unrelated controls. These included enhanced neural progenitor proliferation but impaired neuronal differentiation, maturation, and migration, and increased endoplasmic reticulum (ER) stress. Both the neuronal migration deficit and elevated ER stress could be selectively rescued by different pharmacologic agents. Neuronal gene expression was also dysregulated in the AP, including reduced expression of genes related to behavior, psychological disorders, neuritogenesis, neuronal migration, and Wnt, axonal guidance, and GABA receptor signaling. The UM model instead exhibited upregulated expression of genes in many of these same pathways, suggesting that molecular compensation could have contributed to the lack of neurodevelopmental phenotypes in this model. However, both AP- and UM-derived neurons exhibited shared alterations of neuronal function, including increased action potential firing and elevated cholinergic activity, consistent with increased homomeric CHRNA7 channel activity.
Conclusions: These data define both diagnosis-associated cellular phenotypes and shared functional anomalies related to CHRNA7 duplication that may contribute to variable phenotypic penetrance in individuals with 15q13.3 duplication. The capacity for pharmacological agents to rescue some neurodevelopmental anomalies associated with diagnosis suggests avenues for intervention for carriers of this duplication and other CNVs that cause related disorders.
Keywords: CHRNA7; Chromosome 15q13.3 duplication; Copy number variants; Cortical neurons; Induced pluripotent stem cells; Neurodevelopmental disorders; Psychiatric disease.
© 2021. The Author(s).
Conflict of interest statement
F. Urano received JTV-519 from the National Center for Advancing Translational Sciences for developing small molecule-based therapies for ER stress-related disorders and shares the intellectual property rights related to JTV-519 with the National Institutes of Health. F. Urano is an inventor of the patent related to ER calcium stabilizers, 10,441,574, B2 TREATMENT FOR WOLFRAM SYNDROME AND OTHER ER STRESS DISORDERS. The other authors declare that they have no competing interests.
Figures

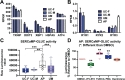
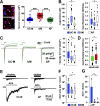
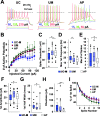
References
Publication types
MeSH terms
Substances
Grants and funding
- R01 NS114551/NS/NINDS NIH HHS/United States
- U54 HD087011/HD/NICHD NIH HHS/United States
- P30 CA091842/CA/NCI NIH HHS/United States
- UL1 TR000448/TR/NCATS NIH HHS/United States
- P60 DK020579/DK/NIDDK NIH HHS/United States
- P30 DK020579/DK/NIDDK NIH HHS/United States
- R01 DK112921/DK/NIDDK NIH HHS/United States
- UL1 TR002345/TR/NCATS NIH HHS/United States
- R01 MH124808/MH/NIMH NIH HHS/United States
- R56 NS114551/NS/NINDS NIH HHS/United States
- P50 HD103525/HD/NICHD NIH HHS/United States
- U01 HG007530/NS/NINDS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

