Lytic archaeal viruses infect abundant primary producers in Earth's crust
- PMID: 34330907
- PMCID: PMC8324899
- DOI: 10.1038/s41467-021-24803-4
Lytic archaeal viruses infect abundant primary producers in Earth's crust
Abstract
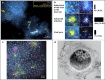
The continental subsurface houses a major portion of life's abundance and diversity, yet little is known about viruses infecting microbes that reside there. Here, we use a combination of metagenomics and virus-targeted direct-geneFISH (virusFISH) to show that highly abundant carbon-fixing organisms of the uncultivated genus Candidatus Altiarchaeum are frequent targets of previously unrecognized viruses in the deep subsurface. Analysis of CRISPR spacer matches display resistances of Ca. Altiarchaea against eight predicted viral clades, which show genomic relatedness across continents but little similarity to previously identified viruses. Based on metagenomic information, we tag and image a putatively viral genome rich in protospacers using fluorescence microscopy. VirusFISH reveals a lytic lifestyle of the respective virus and challenges previous predictions that lysogeny prevails as the dominant viral lifestyle in the subsurface. CRISPR development over time and imaging of 18 samples from one subsurface ecosystem suggest a sophisticated interplay of viral diversification and adapting CRISPR-mediated resistances of Ca. Altiarchaeum. We conclude that infections of primary producers with lytic viruses followed by cell lysis potentially jump-start heterotrophic carbon cycling in these subsurface ecosystems.
© 2021. The Author(s).
Conflict of interest statement
The author declares no competing interests.
Figures




References
-
- Magnabosco C, et al. The biomass and biodiversity of the continental subsurface. Nat. Geosci. 2018;11:707–717. doi: 10.1038/s41561-018-0221-6. - DOI

