Exploring chromatin structural roles of non-coding RNAs at imprinted domains
- PMID: 34338292
- PMCID: PMC8421051
- DOI: 10.1042/BST20210758
Exploring chromatin structural roles of non-coding RNAs at imprinted domains
Abstract
Different classes of non-coding RNA (ncRNA) influence the organization of chromatin. Imprinted gene domains constitute a paradigm for exploring functional long ncRNAs (lncRNAs). Almost all express an lncRNA in a parent-of-origin dependent manner. The mono-allelic expression of these lncRNAs represses close by and distant protein-coding genes, through diverse mechanisms. Some control genes on other chromosomes as well. Interestingly, several imprinted chromosomal domains show a developmentally regulated, chromatin-based mechanism of imprinting with apparent similarities to X-chromosome inactivation. At these domains, the mono-allelic lncRNAs show a relatively stable, focal accumulation in cis. This facilitates the recruitment of Polycomb repressive complexes, lysine methyltranferases and other nuclear proteins - in part through direct RNA-protein interactions. Recent chromosome conformation capture and microscopy studies indicate that the focal aggregation of lncRNA and interacting proteins could play an architectural role as well, and correlates with close positioning of target genes. Higher-order chromatin structure is strongly influenced by CTCF/cohesin complexes, whose allelic association patterns and actions may be influenced by lncRNAs as well. Here, we review the gene-repressive roles of imprinted non-coding RNAs, particularly of lncRNAs, and discuss emerging links with chromatin architecture.
Keywords: CTCF; chromatin; epigenetics; genomic imprinting; non-coding RNA.
© 2021 The Author(s).
Conflict of interest statement
The authors declare that there are no competing interests associated with the manuscript.
Figures
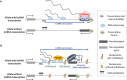
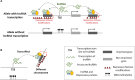
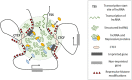
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

