Dynamics of SARS-CoV-2 Spike Proteins in Cell Entry: Control Elements in the Amino-Terminal Domains
- PMID: 34340537
- PMCID: PMC8406164
- DOI: 10.1128/mBio.01590-21
Dynamics of SARS-CoV-2 Spike Proteins in Cell Entry: Control Elements in the Amino-Terminal Domains
Abstract
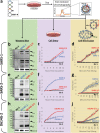
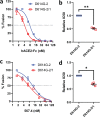
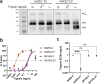
Selective pressures drive adaptive changes in the coronavirus spike proteins directing virus-cell entry. These changes are concentrated in the amino-terminal domains (NTDs) and the receptor-binding domains (RBDs) of complex modular spike protein trimers. The impact of this hypervariability on virus entry is often unclear, particularly with respect to sarbecovirus NTD variations. Therefore, we constructed indels and substitutions within hypervariable NTD regions and used severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus-like particles and quantitative virus-cell entry assays to elucidate spike structures controlling this initial infection stage. We identified NTD variations that increased SARS-CoV-2 spike protein-mediated membrane fusion and cell entry. Increased cell entry correlated with greater presentation of RBDs to ACE2 receptors. This revealed a significant allosteric effect, in that changes within the NTDs can orient RBDs for effective virus-cell binding. Yet, those NTD changes elevating receptor binding and membrane fusion also reduced interdomain associations, leaving spikes on virus-like particles susceptible to irreversible inactivation. These findings parallel those obtained decades ago, in which comparisons of murine coronavirus spike protein variants established inverse relationships between membrane fusion potential and virus stability. Considerable hypervariability in the SARS-CoV-2 spike protein NTDs also appear to be driven by counterbalancing pressures for effective virus-cell entry and durable extracellular virus infectivity. These forces may selectively amplify SARS-CoV-2 variants of concern. IMPORTANCE Adaptive changes that increase SARS-CoV-2 transmissibility may expand and prolong the coronavirus disease 2019 (COVID-19) pandemic. Transmission requires metastable and dynamic spike proteins that bind viruses to cells and catalyze virus-cell membrane fusion. Using newly developed assays reflecting these two essential steps in virus-cell entry, we focused on adaptive changes in SARS-CoV-2 spike proteins and found that deletions in amino-terminal domains reset spike protein metastability, rendering viruses less stable yet more poised to respond to cellular factors that prompt entry and subsequent infection. The results identify adjustable control features that balance extracellular virus stability with facile virus dynamics during cell entry. These equilibrating elements warrant attention when monitoring the evolution of pandemic coronaviruses.
Keywords: SARS-CoV-2; coronavirus; coronavirus spike protein; membrane fusion; virus entry; virus receptors.
Figures


References
-
- Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, Chen HD, Chen J, Luo Y, Guo H, Jiang RD, Liu MQ, Chen Y, Shen XR, Wang X, Zheng XS, Zhao K, Chen QJ, Deng F, Liu LL, Yan B, Zhan FX, Wang YY, Xiao GF, Shi ZL. 2020. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. doi:10.1038/s41586-020-2012-7. - DOI - PMC - PubMed
-
- Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, Hengartner N, Giorgi EE, Bhattacharya T, Foley B, Hastie KM, Parker MD, Partridge DG, Evans CM, Freeman TM, de Silva TI, Sheffield C-GG, McDanal C, Perez LG, Tang H, Moon-Walker A, Whelan SP, LaBranche CC, Saphire EO, Montefiori DC, Sheffield COVID-19 Genomics Group . 2020. Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell 182:812–827.e19. doi:10.1016/j.cell.2020.06.043. - DOI - PMC - PubMed
-
- Zhang L, Jackson CB, Mou H, Ojha A, Peng H, Quinlan BD, Rangarajan ES, Pan A, Vanderheiden A, Suthar MS, Li W, Izard T, Rader C, Farzan M, Choe H. 2020. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat Commun 11:6013. doi:10.1038/s41467-020-19808-4. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
